| One tool to find them all: A case of data integration and querying in a distributed LIMS platform 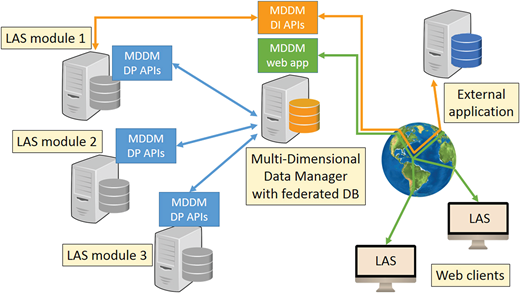 In this early 2019 journal article published in the journal Database, Grand et al. of the Candiolo Cancer Institute present the fine details of their laboratory information management system (LIMS) Laboratory Assistant Suite (LAS) for cancer and other genomic research. Citing "a substantial mismatch between the LIMS solutions on offer and the functional requirements dictated by research practice," the authors describe the requirements they had for a LIMS in their institution and how they went about creating it. After describing the data models, functionalities, modular architecture, and its usage, the authors conclude that their LAS, in conjunction with a custom data management module, allows researchers to "execute complex queries without any knowledge of query languages or database structures, and easily integrate heterogeneous data stored in multiple databases," while also resulting in an improvement in data quality, a reduction in data entry and retrieval, and new insights with the enabled data interconnections. In this early 2019 journal article published in the journal Database, Grand et al. of the Candiolo Cancer Institute present the fine details of their laboratory information management system (LIMS) Laboratory Assistant Suite (LAS) for cancer and other genomic research. Citing "a substantial mismatch between the LIMS solutions on offer and the functional requirements dictated by research practice," the authors describe the requirements they had for a LIMS in their institution and how they went about creating it. After describing the data models, functionalities, modular architecture, and its usage, the authors conclude that their LAS, in conjunction with a custom data management module, allows researchers to "execute complex queries without any knowledge of query languages or database structures, and easily integrate heterogeneous data stored in multiple databases," while also resulting in an improvement in data quality, a reduction in data entry and retrieval, and new insights with the enabled data interconnections.
From command-line bioinformatics to bioGUI 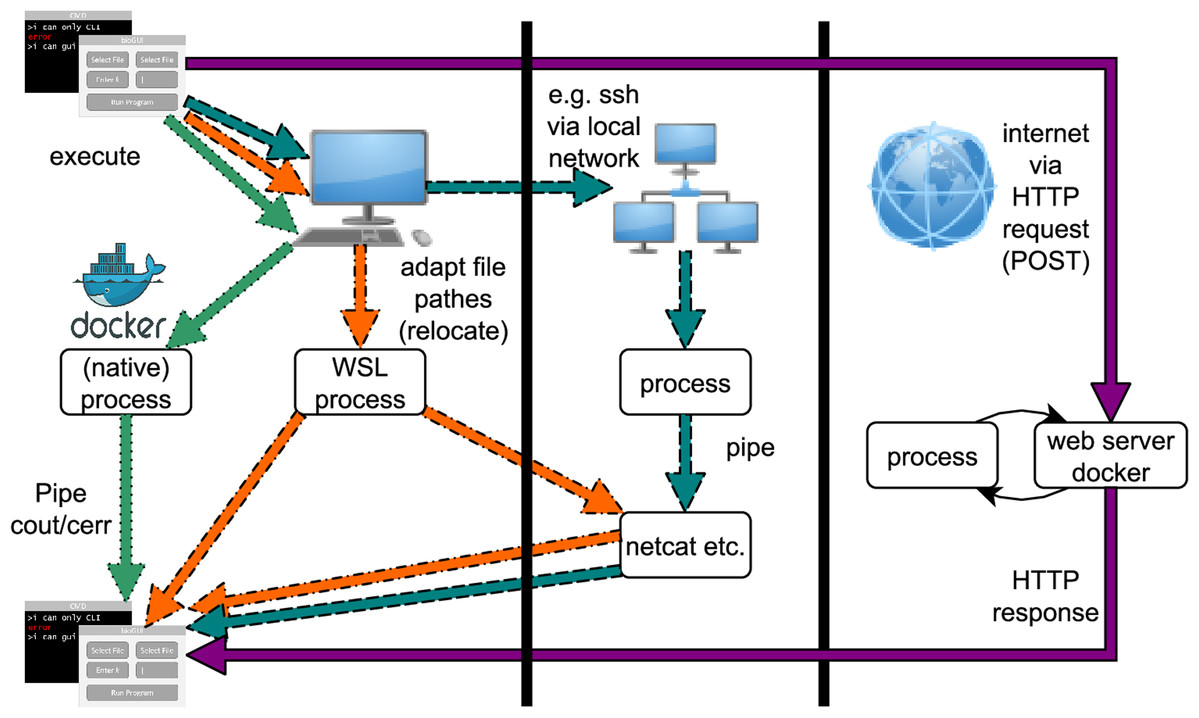 The topic of making bioinformatics applications more approachable to researchers and students has been discussed off and on for years, and some efforts have even been made in that regard. Another step forward for bioinformatics applications is offered by Joppich and Zimmer of Ludwig-Maximilians-Universität München, with their open-source bioGUI. The software attempts to address two problems of bioinformatics applications that rely heavily on the command line: many of them work on Unix-based systems but not Microsoft Windows, and researchers have a tendency to shy away from complex command-line apps despite their utility. The authors present in detail their framework and its use cases, showing how a graphical user interface or GUI can make many such command-line apps more approachable. They conclude that providing a GUI and easy-to-use install modules for bioinformatics apps using the command line makes "execution and usage of these tools more comfortable" while allowing scientists to better analyze their data. The topic of making bioinformatics applications more approachable to researchers and students has been discussed off and on for years, and some efforts have even been made in that regard. Another step forward for bioinformatics applications is offered by Joppich and Zimmer of Ludwig-Maximilians-Universität München, with their open-source bioGUI. The software attempts to address two problems of bioinformatics applications that rely heavily on the command line: many of them work on Unix-based systems but not Microsoft Windows, and researchers have a tendency to shy away from complex command-line apps despite their utility. The authors present in detail their framework and its use cases, showing how a graphical user interface or GUI can make many such command-line apps more approachable. They conclude that providing a GUI and easy-to-use install modules for bioinformatics apps using the command line makes "execution and usage of these tools more comfortable" while allowing scientists to better analyze their data. |