Posted on January 30, 2024
By LabLynx
Journal articles
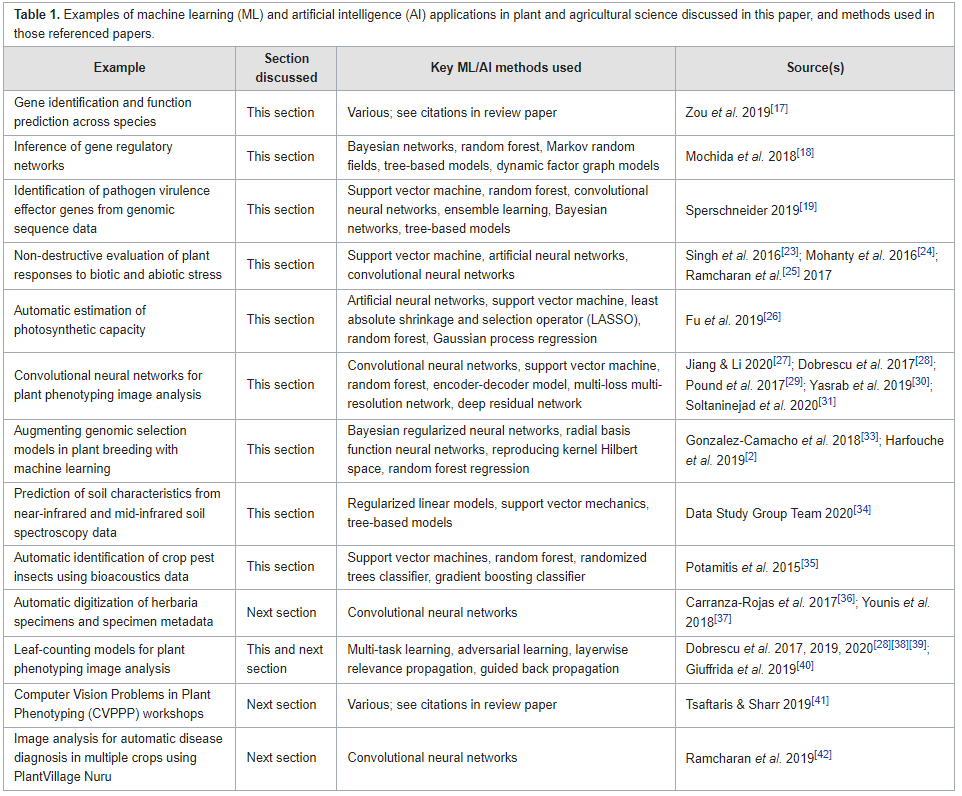
In this 2023 article published in the journal
F1000Research, Williamson
et al. identify and discuss "eight key challenges in data management that must be addressed to further unlock the potential of [artificial intelligence] in crop and agronomic research." Noting the state of the agricultural research landscape and growing potentials of artificial intelligence and machine learning in the field, the authors perform a literature review that better shapes the nuances of those eight key challenges: data heterogeneity, data selection and digitization, data linkages, standardization and curation of data, sufficient training and ground truth data, system access, data access, and engagement. After examining these challenges in detail, the authors conclude there's a definitive "need for a more systemic change in how research in this domain is conducted, incentivized, supported, and regulated," with aspects such as stronger collaboration, more efficient machine learning methods, improved data curation, and improved data management methods.
Posted on January 23, 2024
By LabLynx
Journal articles
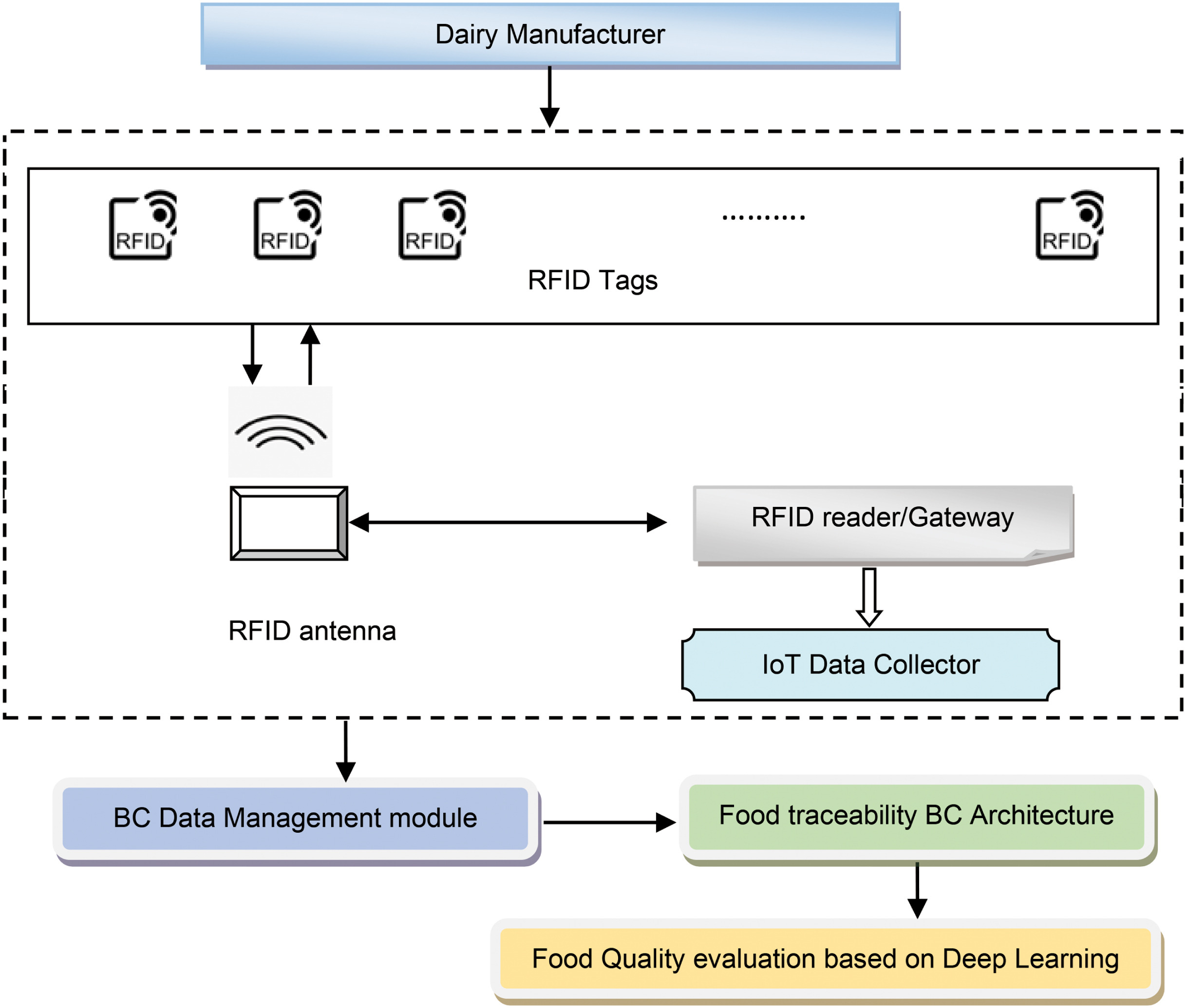
In this 2023 paper published in the journal
High-Confidence Computing, Manisha and Jagadeeshwar present their custom food quality traceability system, which combines various tools like blockchain, internet of things (IoT) mechanisms, and deep learning architecture for improving traceability about perishable food supply chains (PFSCs). Noting previous works that incorporated some but not all of these elements, each with their own downsides, the authors chose a system model that incorporates blockchain-enabled RFID scans that, upon verification, get added to the overall secure ledger, along with associated IoT-base metadata from sensors gauging humidity and temperature. The duo turns to milk manufacturing and distribution in their case study, applied to their blockchain-enabled deep residual network (BC-DRN) methodology. After examining and comparing their results to other prevalent methodologies used in supply chain management (SCM). The authors conclude that their BC-DRN traceability system, when gauged on metrics like sensitivity, response time, and testing accuracy, beat out other methods. They add "the performance of the devised scheme can be improved by considering better feature extraction techniques. "
Posted on January 16, 2024
By LabLynx
Journal articles
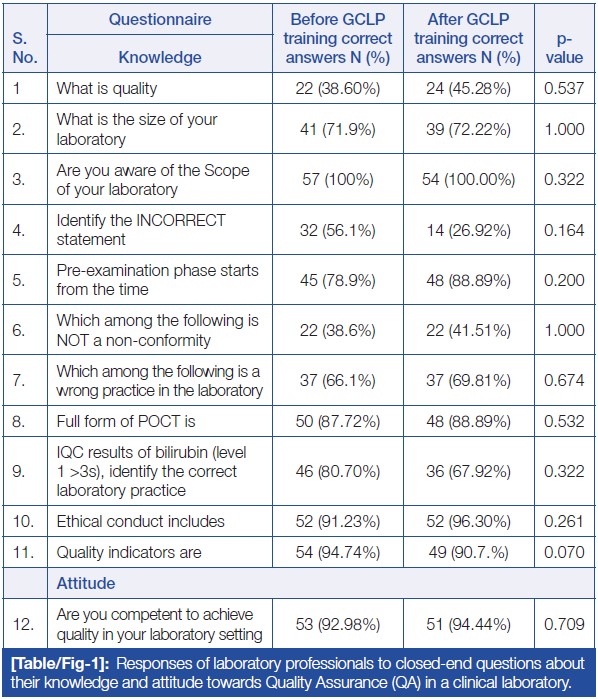
In this brief 2023 article published in
Journal of Clinical and Diagnostic Research, Patel
et al. provide the results of their survey-based analysis of laboratorians' knowledge, attitude, and practice (KAP) towards laboratory quality through good clinical laboratory practice (GCLP) training. Noting the "ethical obligation to provide accurate and precise results that are cost- and time-effective," the authors state that it's imperative for clinical laboratory personnel to adhere to quality planning and system implementation, while also possessing an understanding of quality management principles as they apply to the lab. The authors describe their survey format and present the results of their survey, concluding that while no statistically significant differences could be found towards staff attitudes towards quality in the lab after GCLP training, laboratorians in their survey acknowledged the benefits of GCLP guidelines and accreditation, as well as the importance of training on such matters. "Such training and assessments would also aid in evaluating the performance of laboratory staff, contributing to improved learning, execution of GLPs, and consistent patient care services."
Posted on January 9, 2024
By LabLynx
Journal articles
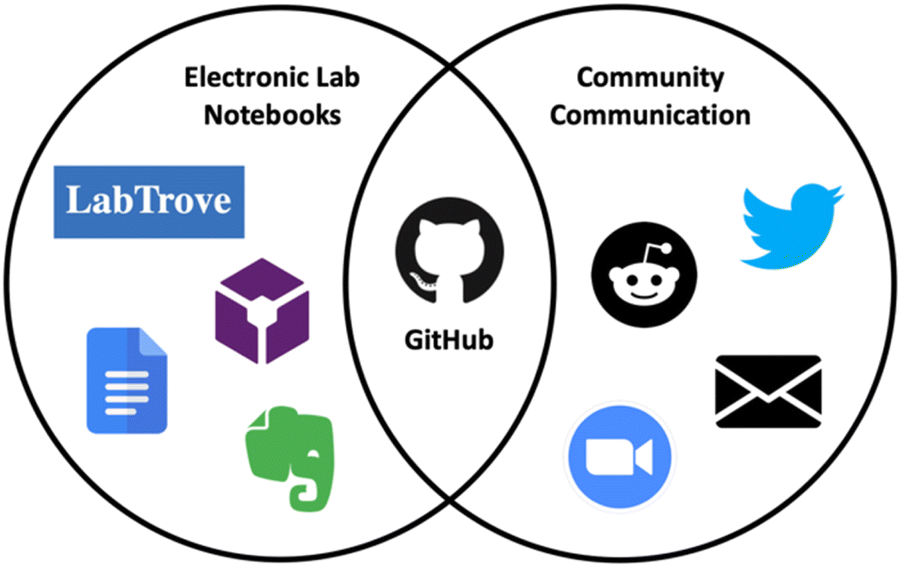
In this 2023 paper published in the journal
Digital Discover, Scroggie
et al. of the University of Sydney present their efforts towards utilizing the open developer platform GitHub as an electronic laboratory notebook (ELN) for chemistry research. Noting a lack of open-source ELNs with a focus on non-organic chemistry that have a wealth of collaboration tools, as well as problems with expensive and inflexible commercial ELNs, the authors turned to the many open facets of GitHub to repurpose its workings for synthetic chemistry projects. After a brief discussion of GitHub, the authors explain how they used the various facets of GitHub for ELN-related tasks, including notebooks, data and metadata management, and collaborative tools. They also acknowledged several shortcomings of their approach, including learning Markdown, dealing with data storage limitations, and integrating discipline-specific applications. The authors conclude that some of GitHub's features "are undeniably more oriented towards coders, such as the Actions tab in which users can set up workflows using code, these features do not detract from GitHub's usefulness as an ELN, which lies mainly in its adaptability and capacity for knowledge-sharing and collaboration."
Posted on January 2, 2024
By LabLynx
Journal articles
In this 2023 paper published in the journal
GigaScience, Nieminen
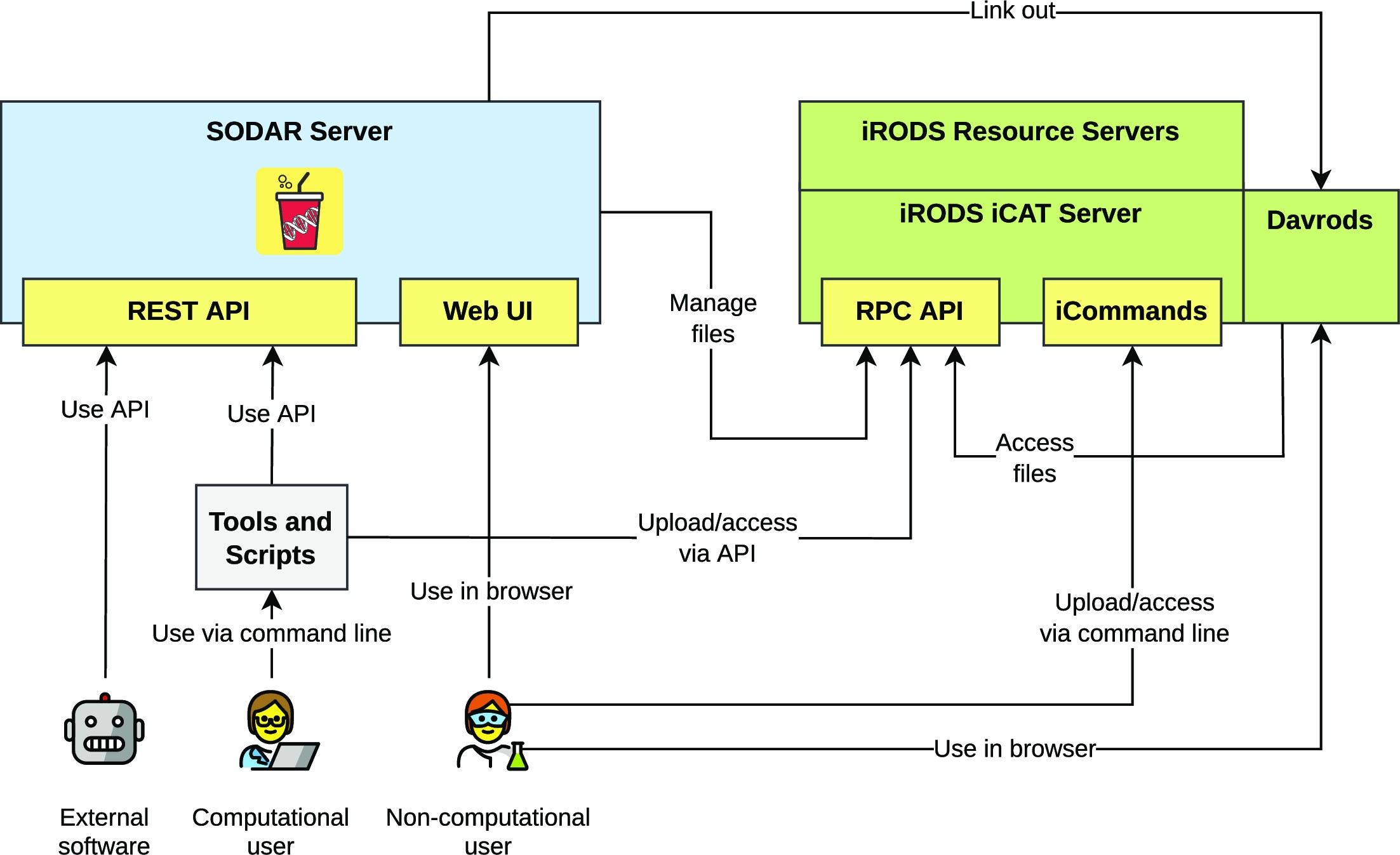
et al. of the Berlin Institute of Health at Charité–Universitätsmedizin Berlin present SODAR (System for Omics Data Access and Retrieval), an open-source scientific data management system (SDMS) with a focus on omics data management during multiassay research studies. Noting numeroud data management challenges and dearth of open-source options, the authors describe the software framework, features, and limitations of the system. Highlighting SODAR's "programmable application programming interfaces (APIs) and command-line access for metadata and file storage," the authors conclude that SODAR can readily "support multiple technologies such as whole genome sequencing, single-cell sequencing, proteomics, and mass spectrometry," though some aspects such as automated data export and "data commons" access are currently not available.
Posted on December 26, 2023
By LabLynx
Journal articles

In this 2023 paper published in
Healthcare Informatics Research, Chang
et al. perform a literature review of current research in order to better characterize the currently viewed benefits of of artificial intelligence (AI), the internet of things (IoT), and personal health records (PHR) in the healthcare setting. Noting "limited empirical evidence regarding the benefits of information technology in healthcare settings," the authors examined 24 reviews and meta-analysis studies on these three technologies, with a strong focus on four outcome domains of clinical, psycho-behavioral, managerial, and socioeconomic implications. After detailing their findings, the authors conclude that "AI and PHRs can enhance clinical outcomes, while IoT holds promise for boosting managerial efficiency," though further research is required to address "the organizational and socioeconomic benefits of PHR," as well as the greater role of IoT in the healthcare setting.
Posted on December 19, 2023
By LabLynx
Journal articles
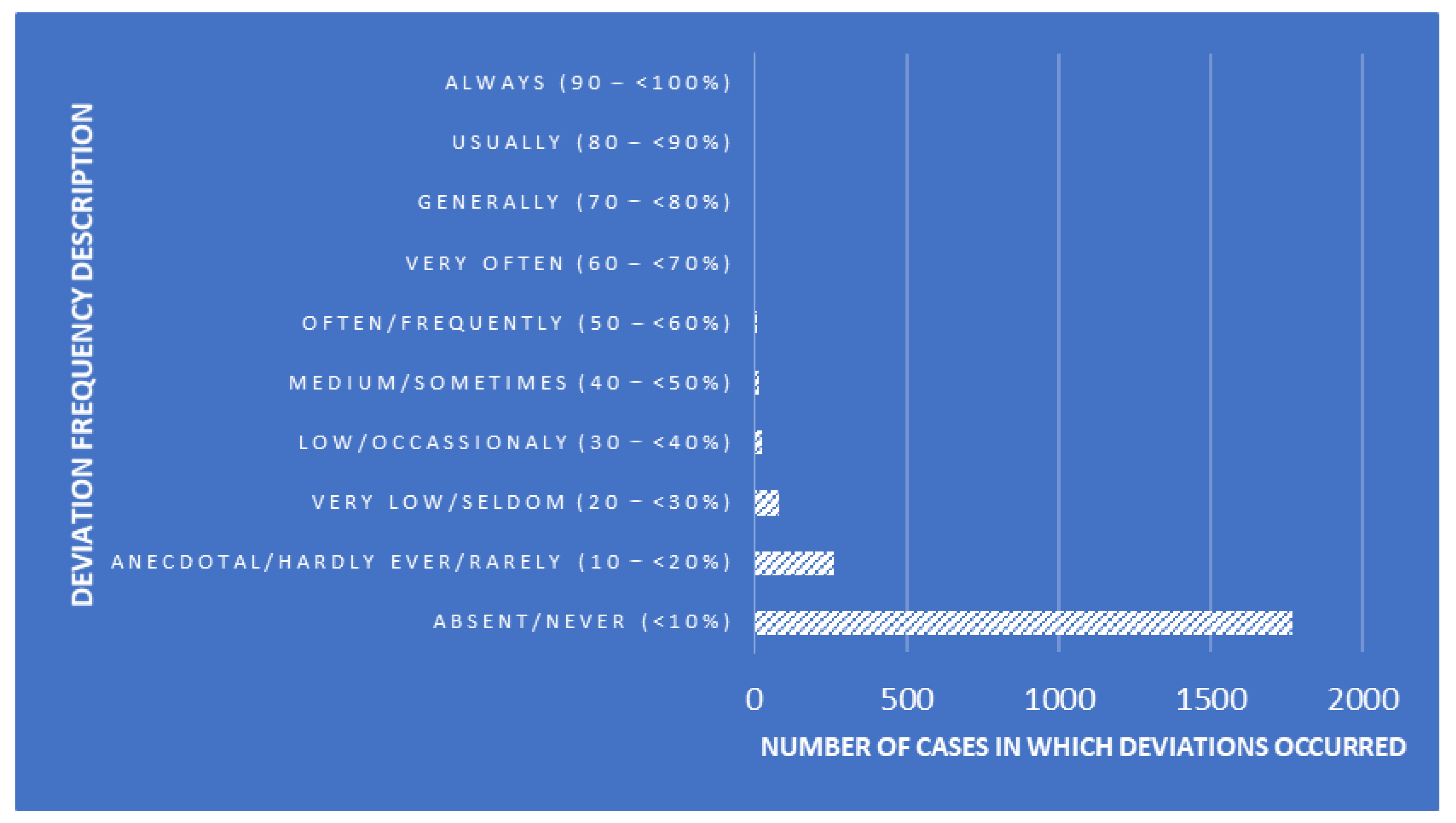
Like many other parts of the food and beverage industry, the meat processing industry is guided by regulations that mandate the safety of the final product to the consumer. This includes laboratory testing for
Trichinella spp., a nematode parasite responsible for the disease trichinosis. In order to ensure timely and accurate results, laboratories in the meat industry must act with purpose, implementing quality assurance (QA) practices that incorporate quality management systems (QMSs). In this 2023 journal article, Villegas-Pérez
et al. examine the state of QA and QMSs in the in-house laboratories of Southern Spain's slaughterhouses and game-handling establishments, using canonical discriminant analysis (CDA) to gauge the effectiveness of those labs' practices. After lengthy review and discussion, the authors conclude that their CDA-based tools were able to discover "deficiencies in processes and procedures, necessitating measures for result reliability, due to facilities’ unfamiliarity with extensive QMS documentation." The authors offer several recommendations to fix those deficiencies.
Posted on December 12, 2023
By LabLynx
Journal articles
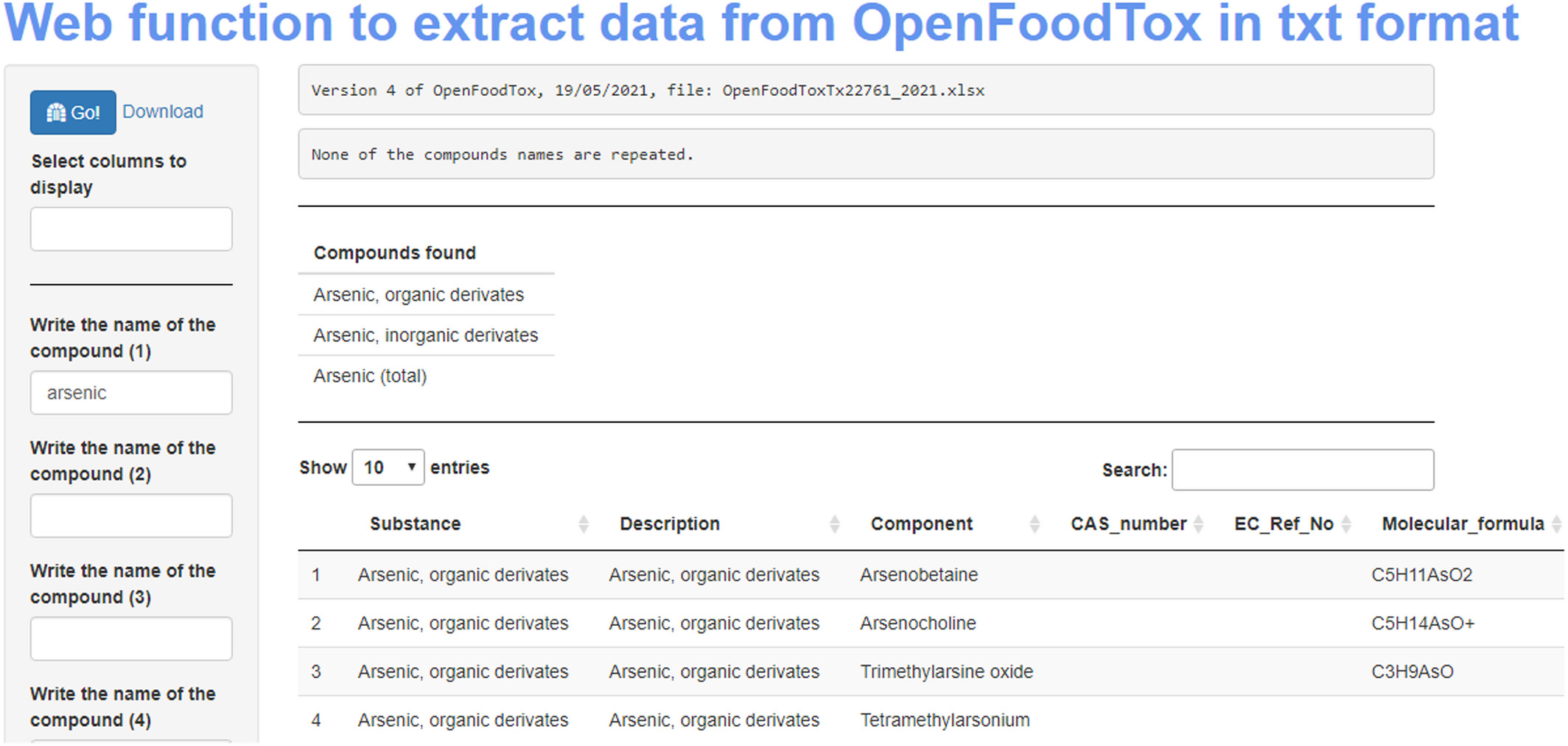
In this brief paper published in the
EFSA Journal (from the European Food Safety Authority), Pineda-Pampliega
et al. of the Norwegian Scientific Committee for Food and Environment (VKM) discuss the specifics of their work-related project to see "if existing commonly used [food safety] databases in risk assessment are in line with the FAIR principles" (which ensure data and information is findable, accessible, interoperable, and reusable), as well as to determine how to improve deficiencies in meeting FAIR principles. The authors describe their approach using data from both the EFSA's Chemical Hazards Database, OpenFoodTox, and the Institute of Marine Research's Seafood database. After evaluating these databases and describing the necessary framework activities to make them more FAIR, the authors conclude that their methods of using the R programming language, Shiny, GitHub,
Zenodo, and appropriate file formats "is an essential step to ensuring the success of the future risk–benefit assessment [towards food safety], by offering more timely results with adequate spending of human and economic resources."
Posted on December 4, 2023
By LabLynx
Journal articles
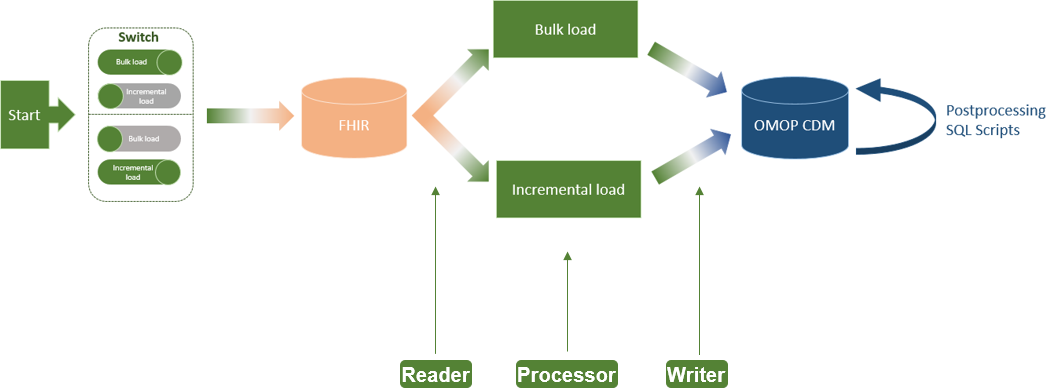
In this 2023 paper published in
JMIR Medical Informatics, Henke
et al. of Technische Universität Dresden present the results of their effort to add "incremental loading" to the Medical Informatics in Research and Care in University Medicine's (MIRACUM's) clinical trial recruitment support systems (CTRSSs). Those CTRSSs already allows bulk loading of German-based FHIR data, supporting "the possibilities for multicentric and even international studies," but MIRACUM needed greater efficiencies when updating such data on a daily, incremental basis. The paper presents their literature review and approach to adding incremental loading to their systems. They conclude that the extract-transfer-load (ETL) "process no longer needs to be executed as a bulk load every day" with the change, instead being able to rely on "using bulk load for an initial load and switching to incremental load for daily updates." They add that this process has international applicability and is not limited to German FHIR data.
Posted on November 28, 2023
By LabLynx
Journal articles
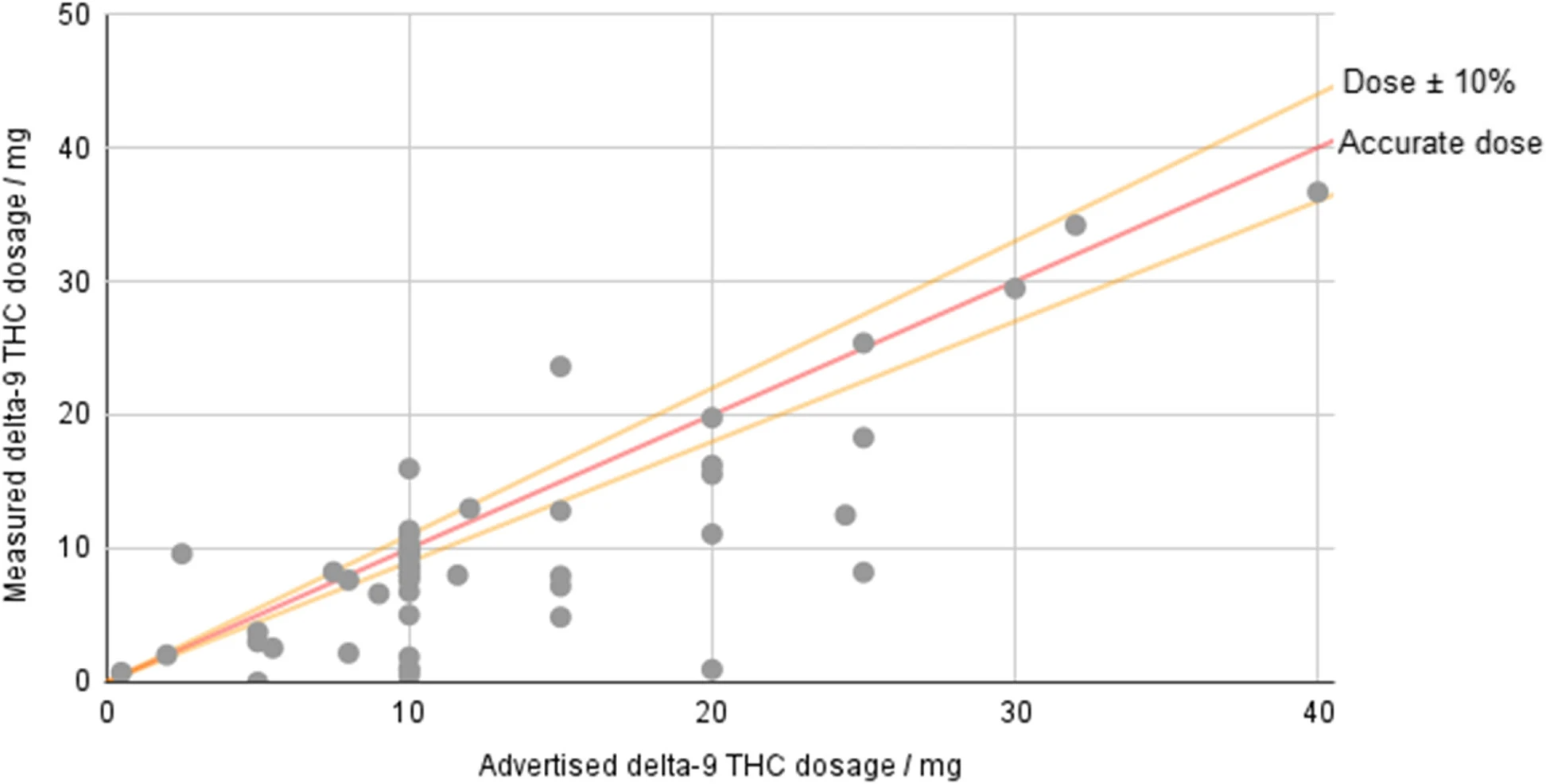
In this 2023 paper published in the
Journal of Cannabis Research, Johnson
et al. examine the current state of hemp-derived delta-9-tetrahydrocannabinol (Δ
9-THC) products on the U.S. market after the passage of the Agriculture Improvement Act of 2018. Noting discrepancies and loopholes in the legislation, the authors performed laboratory analyses on 53 hemp-derived Δ
9-THC products from 48 brands, while also examining aspects such as age verification, labeling consistency, and comparison of reported company values vs. analyzed values. After describing their methodology and results, the authors conclude that "the legal status of hemp-derived Δ
9-THC products in America essentially permits their open sale while placing very few requirements on the companies selling them." The end result includes finding, for example, products "that have 3.7 times the THC content of edibles in adult-use states," as well as inaccurately labeled products.
Posted on November 20, 2023
By LabLynx
Journal articles

"Artificial intelligence" (AI) may seem like a buzzword akin to the "nanotechnology" craze of the 2000s, but it is inevitably finding its way into scientific applications, including in the materials sciences. In this December 2022 article published in
npj Computational Materials, Sbailò
et al. present their AI-driven Novel Materials Discovery (NOMAD) toolkit as an extension of their NOMAD Repository & Archive, focused on making materials science data FAIR (findable, accessible, interoperable, and reusable), as well as AI-ready. After introducing the details of their workspace and goals towards adding notebook-based tools to the NOMAD Repository & Archive, the authors dive into the details of NOMAD AI Toolkit. They close by noting their toolkit "offers a selection of notebooks demonstrating such [AI-based] workflows, so that users can understand step by step what was done in publications and readily modify and adapt the workflows to their own needs." They add that the system "will allow for enhanced reproducibility of data-driven materials science papers and dampen the learning curve for newcomers to the field."
Posted on November 13, 2023
By LabLynx
Journal articles
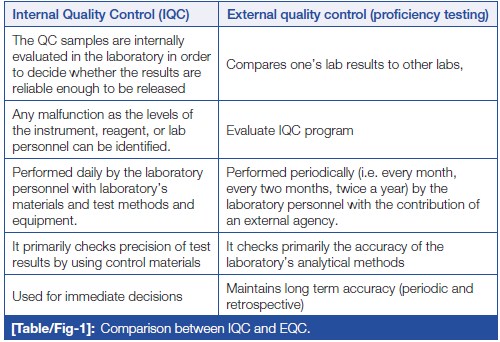

In this 2023 article published in
Journal of Clinical and Diagnostic Research, Naphade
et al. provide a brief introductory-level review of the importance of quality control (QC) to the clinical laboratory. After a brief introduction on clinical lab testing, the authors analyze the wide variety of sources for laboratory errors, covering the pre-analytical, analytical. and post-analytical phases. They then introduce the concept of quality control, followed by explaining how QC is implemented in the laboratory, including through the use of QC materials, statistical control charts, and shifts and trends. They conclude this review by stating that "reliable and confident laboratory testing avoids misdiagnosis, delayed treatment, and unnecessary costing of repeat testing." They add that given these benefits, "the individual laboratory should assess and analyze their own QC process to find out the possible root cause of any digressive test results which are not correlating with patients' clinical presentation or expected response to treatment."
Posted on November 7, 2023
By LabLynx
Journal articles
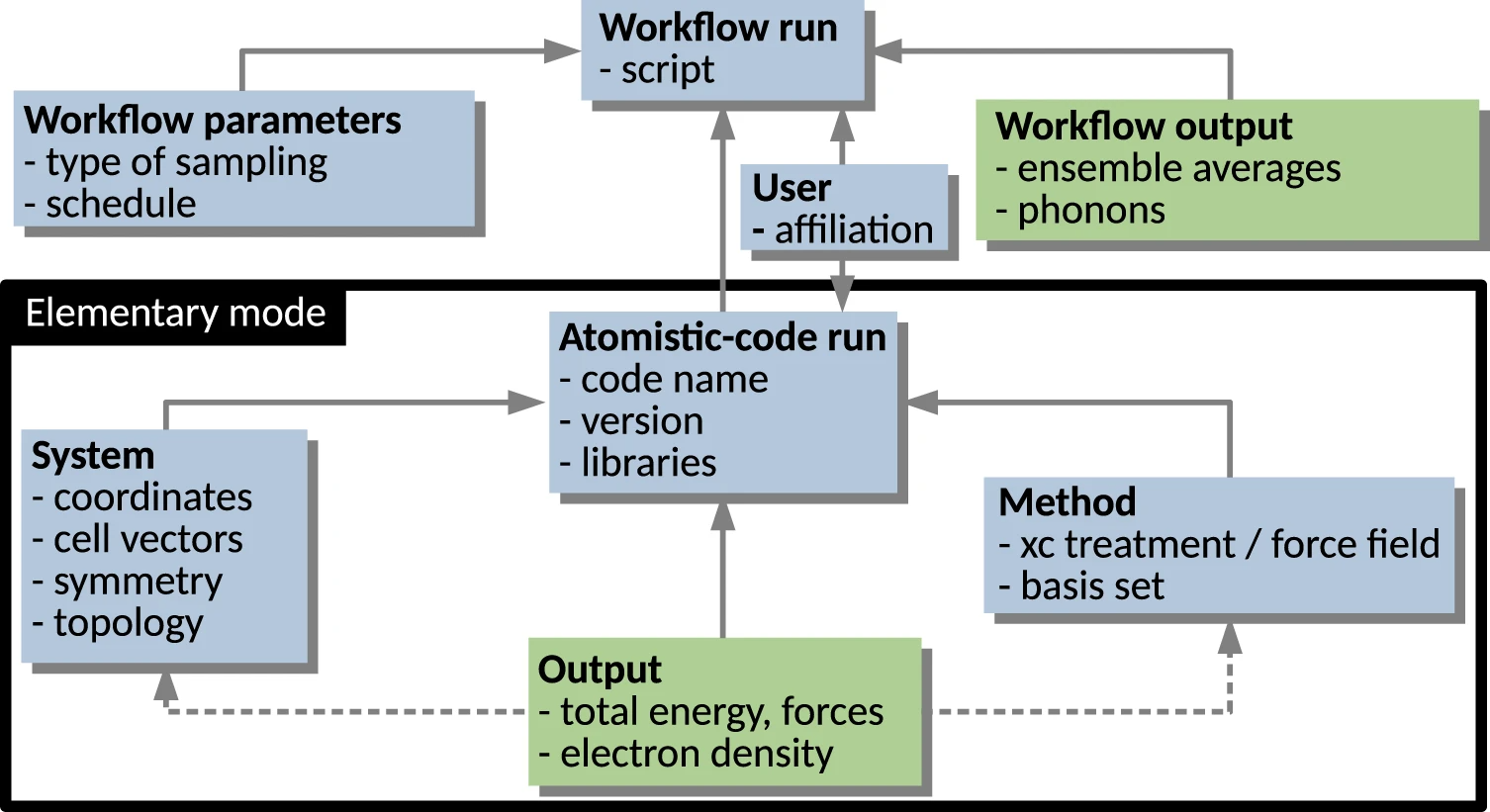
In this 2023 paper published in the journal
Scientific Data, Ghiringhelli
et al. present the results of an international workshop of materials scientists who came together to discuss metadata and data formats as they relate to data sharing in materials science. With a focus on the FAIR principles (findable, accessible, interoperable, and reusable), the authors introduce the concept of materials science data management needs, and the value of metadata towards making shareable data FAIR. They then go into specific use cases, such as electronic-structure calculations, potential-energy sampling, and managing metadata for computational workflows. They also address the nuances of file formats and metadata schemas as they relate to those file formats. They clise with an outlook on the use of ontologies in materials science, followed by conclusions, with a heavy emphasis on "the importance of developing a hierarchical and modular metadata schema in order to represent the complexity of materials science data and allow for access, reproduction, and repurposing of data, from single-structure calculations to complex workflows." They add: "the biggest benefit of meeting the interoperability challenge will be to allow for routine comparisons between computational evaluations and experimental observations."
Posted on October 31, 2023
By Shawn Douglas
Journal articles
In this 2023 journal article published in
International Journal of Molecular Sciences, Jadhav
et al. recommend a more comprehensive approach to analyzing the
Cannabis plant, its extracts, and its constituents, turning to the still-evolving concept of authentomics—"which involves combining the non-targeted analysis of new samples with big data comparisons to authenticated historic datasets"—of the food industry as a paradigm worth shifting towards. After an introduction on metabolomics and a review of its various technologies, the authors examine the current state of cannabis laboratory testing, how its regulated and standardized, and a variety of issues that make cannabis testing less consistent, as well as how cannabis can become adulterated in the grow chain. They then discuss authentomics and apply it to the cannabis industry, as well address what considerations need to be made in the future to make the most of authentomics. The author conclude that an authentomics approach "provides a robust method for verifying the quality of cannabis products."
Posted on October 24, 2023
By LabLynx
Journal articles
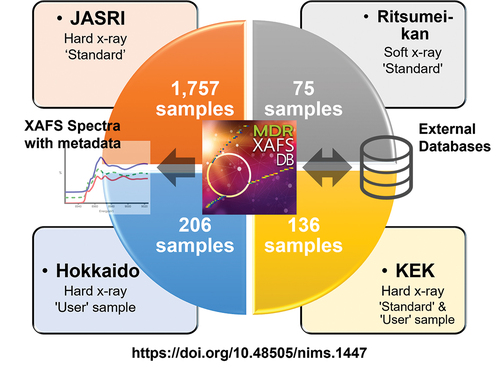
In this 2023 article published in
Science and Technology of Advanced Materials: Methods, Ishii
et al. describe their cross-institutional efforts towards integrating X-ray absorption fine structure (XAFS) spectra and associated metadata across multiple databases to improve materials informatics processes and research methods of materials discovery and analysis. Their new public database, MDR XAFS DB, was developed towards this goal, while addressing two main issues with unifying such data: the difficulties of "designing and collecting metadata describing spectra and sample details, and unifying the vocabulary used in the metadata, including not only metadata items (keys) but also descriptions (values)." After discussing the system's construction and contents, the authors conclude their system "has achieved seamless cross searchability with the use of sample nomenclature so that database users do not have to be aware of the differences in the local metadata of the facilities that provide the data." Though still with some challenges to address, the authors add that while the culture of open data hasn't truly taken hold in materials science, they hope "that this initiative will be a trigger to promote the utilization of materials data."
Posted on October 17, 2023
By LabLynx
Journal articles
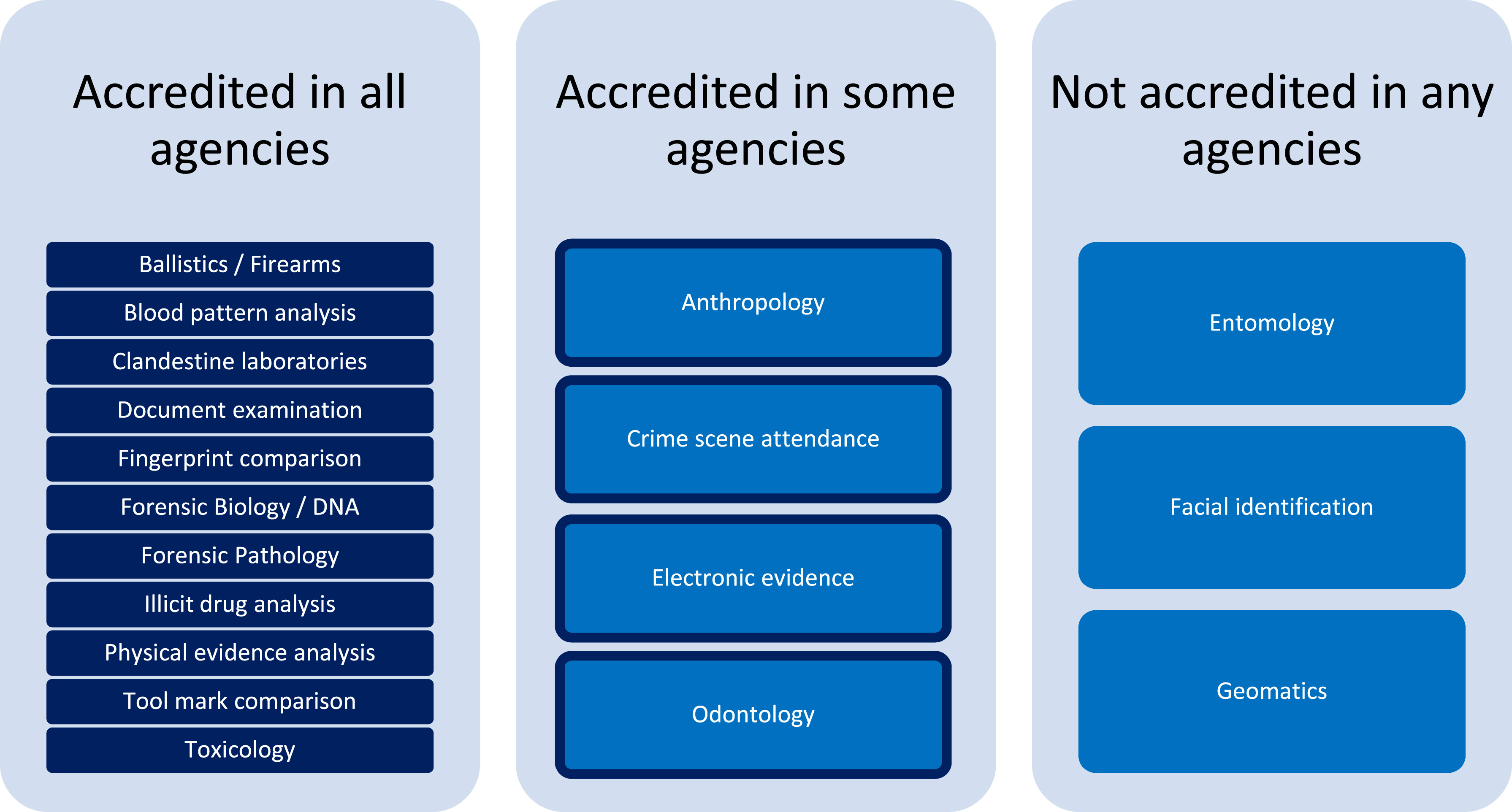
Forensic laboratories are largely bound to performing their actions with quality highly in mind. In fact, standards like ISO/IEC 17025 help such labs focus more on quality with the help of a quality management system (QMS). But even with a strong focus on quality, inconsistencies in recording, managing, and reporting quality issues can be problematic. To that point, Heavey
et al. examine government-based forensic service provider agencies in Australia and New Zealand in this 2023 journal article, using the survey format to gain insight into potential QMS weaknesses. After thoroughly reviewing the methodology and results for their survey, the authors conclude that "the need for further research into the standardization of systems underpinning the management of quality issues in forensic science" is evident from their research, with a greater need "to support transparent and reliable justice outcomes" through improving "evidence-based standard terminology for quality issues in forensic science to support data sharing and reporting, enhance understanding of quality issues, and promote transparency in forensic science."
Posted on October 9, 2023
By LabLynx
Journal articles
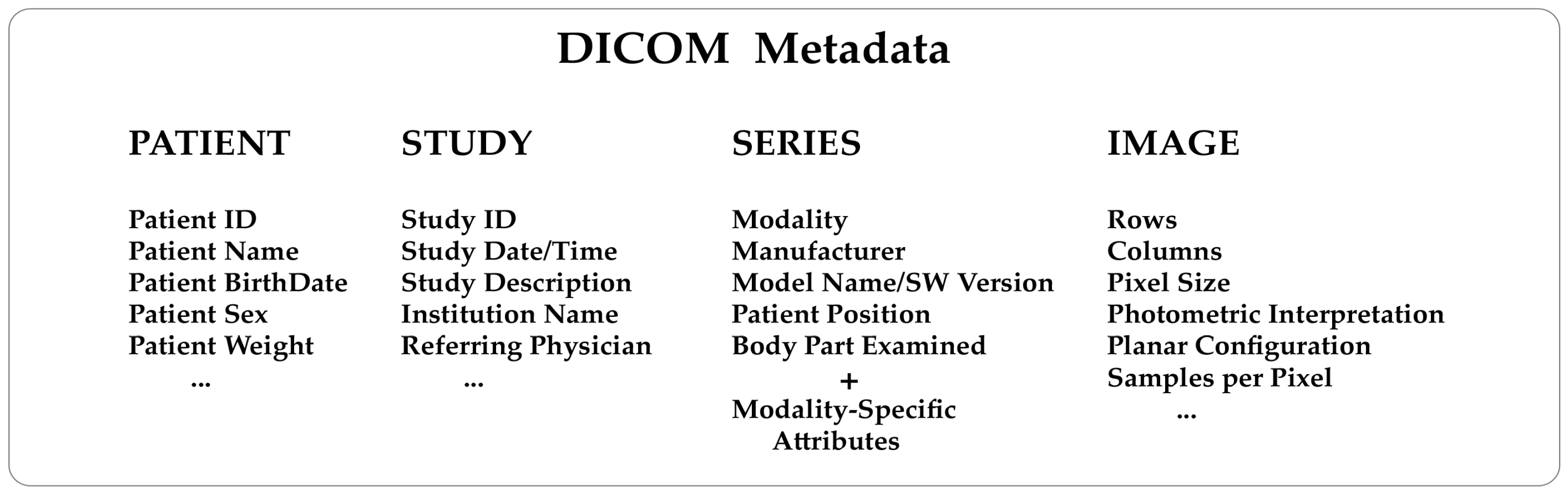

In this brief 2023 article published in the journal
Tomography, Michele Larobina of Consiglio Nazionale delle Ricerche provides background on "the innovation, advantages, and limitations of adopting DICOM and its possible future directions." After an introduction on the Digital Imaging and Communications in Medicine standard, Larobina then emphasizes the benefits of the standard's approach to handling image metadata, followed by the exchange component of the standard. Larobina then examines the strengths and weaknesses of the DICOM standard, before concluding that DICOM has not only demonstrated "the positive influence and added value that a standard can have in a specific field," but also "encouraged and facilitated data exchange between researchers, creating added value for research." Despite this, "the time is ripe to review some of the initial directives" of DICOM, says Larobina, in order for the the standard to remain relevant going forward.
Posted on October 2, 2023
By Shawn Douglas
Journal articles
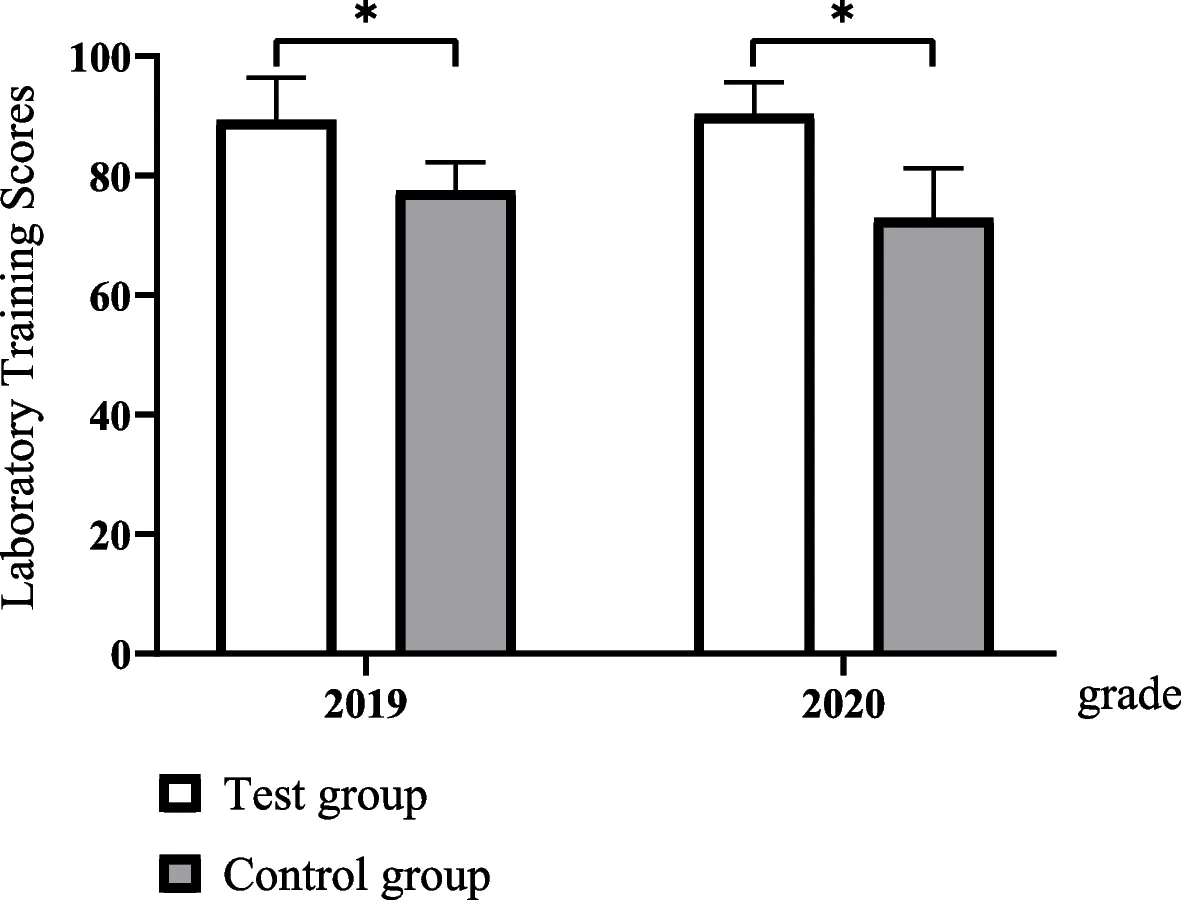
In this article published in
BMC Medical Education, Xu
et al. present their four-stage clinical biochemistry teaching methodology which incorporates elements of the International Organization for Standardization's ISO 15189 standard. Noting a lack of student awareness of ISO 15189
Medical laboratories — Requirements for quality and competence and the value its emphasis on quality management brings to laboratories, the authors set upon incorporating the standard into a case-based learning (CBL) approach. After the authors describe the details of their approach and the survey-based results analysis of its outcomes, the authors conclude that outside of improving student outcomes in clinical biochemistry, "ISO 15189 concepts of continuous improvement were implanted, thereby putting the concept of plan–do–check–act (PDCA) into practice, developing recording habits, and improving communication skills."
Posted on September 26, 2023
By Shawn Douglas
Journal articles
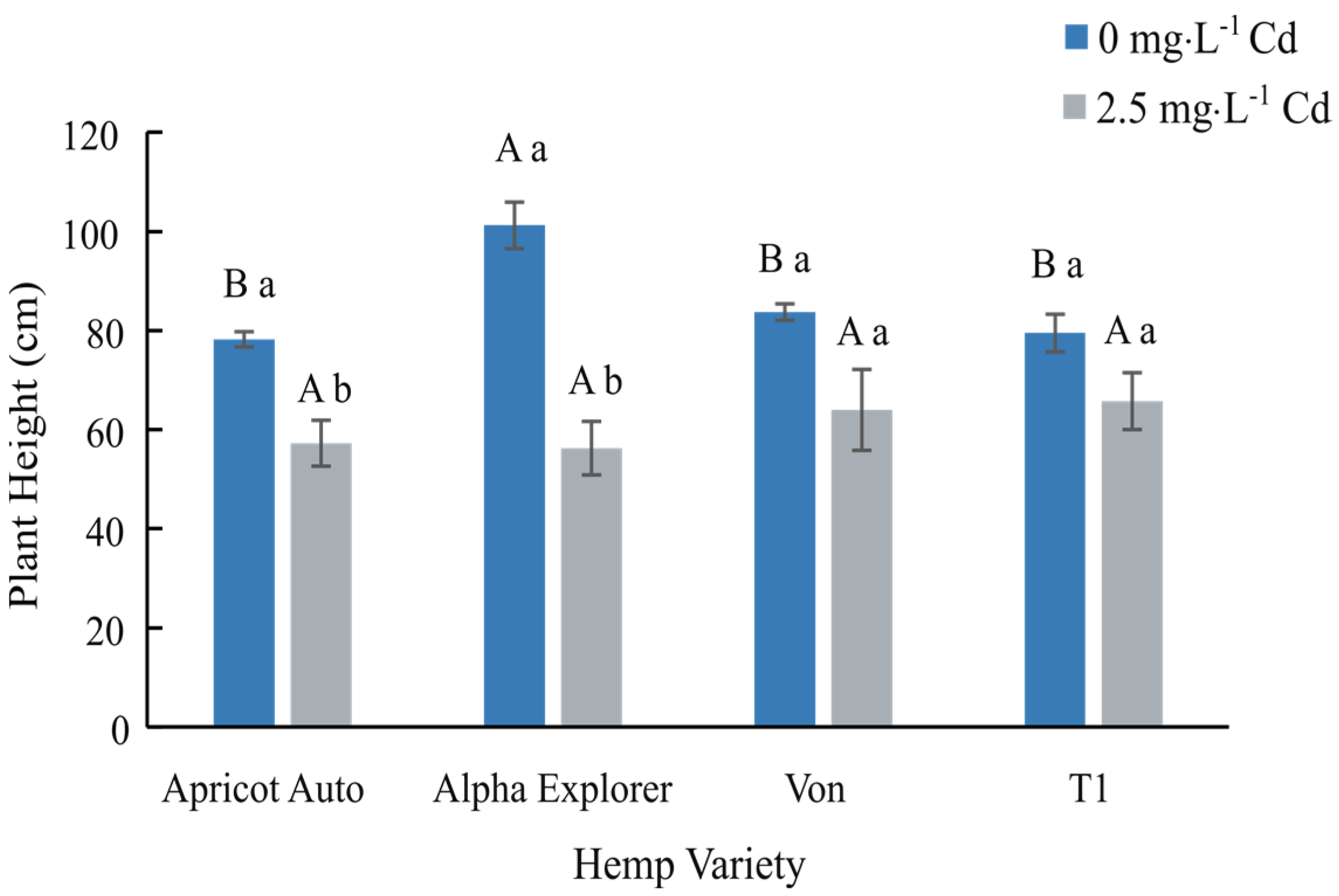
In this 2023 article published in the journal
Water, Marebesi
et al. of the University of Georgia present their findings on phytoremediation of Cadmium (Cd)-contaminated soils using day-neutral and photoperiod-sensitive hemp plants. After a thorough introduction on the topic of using
Cannabis plants for phytoremediation (including the flowers), the authors present their materials and methods, focusing on Apricot Auto, Alpha Explorer, Von, and T1 varieties of hemp. They then present the results of their experiments, addressing plant height, biomass yield, Cd concentrations in various plant tissues, nutrient partitioning, and THC/CBD concentrations. They conclude noting that "while Cd concentration was significantly higher in roots, all four varieties were efficient in translocating Cd from roots to shoots." They also conclude that THC/CBD loss is affected by remediation in some cases, as was height and biomass. Finally, they highlight the importance of Cd testing of flowers for medical markets where the plant is used.
Posted on September 18, 2023
By LabLynx
Journal articles
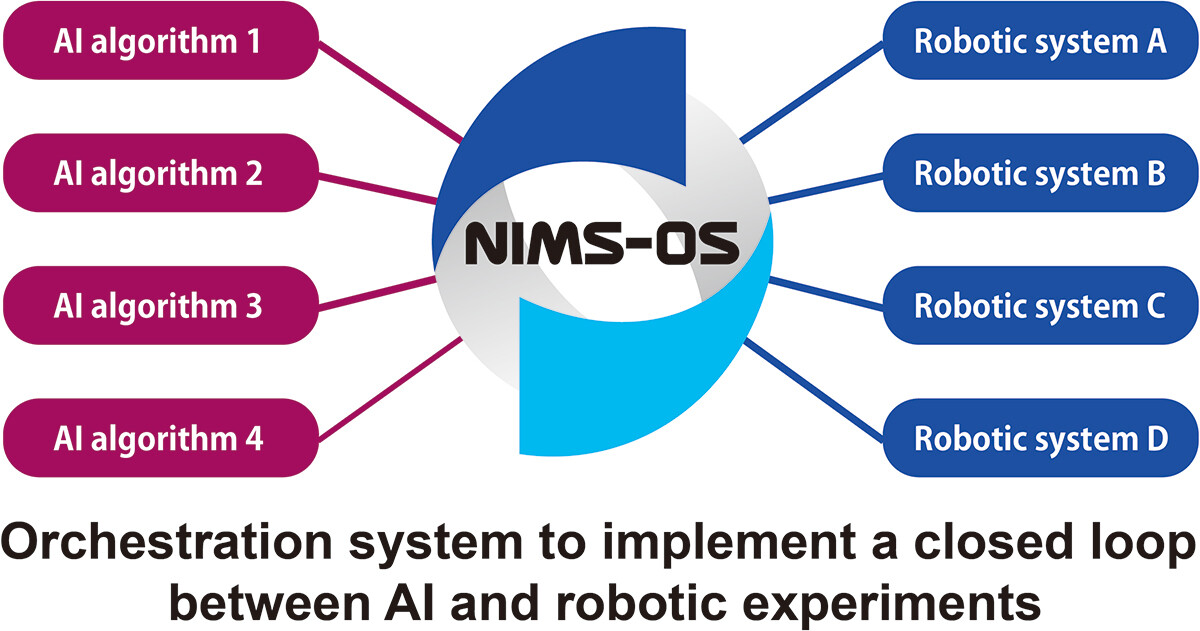
As technological, cultural, and environmental shifts continue to put pressure on the ways academic and private research is conducted, researchers must further strive to get the most out of their work. This holds true across a broad swath of scientific fields, including materials science. Noting the continuing expansion of laboratory automation and software-based tools like artificial intelligence (AI) and machine learning (ML), Tamura
et al. emphasize that materials science and novel materials research can benefit greatly from adopting these tools. In this 2023 paper, the authors present their take on this trend, in the form of NIMS-OS, a Python-based system developed to take advantage of completely automated material discovery experiments. After presenting an introduction on the topic, the authors discuss the development philosophy and approach to making NIMS-OS, while also describing use of both the Python command-line tool and the graphical user interface (GUI)-driven version of the software. The authors conclude that their system demonstrates how "various problems for automated materials exploration to be commonly performed," while at the same time being "designed to be easily adaptable to a variety of original robotic systems in materials science."
 In this 2023 article published in the journal F1000Research, Williamson et al. identify and discuss "eight key challenges in data management that must be addressed to further unlock the potential of [artificial intelligence] in crop and agronomic research." Noting the state of the agricultural research landscape and growing potentials of artificial intelligence and machine learning in the field, the authors perform a literature review that better shapes the nuances of those eight key challenges: data heterogeneity, data selection and digitization, data linkages, standardization and curation of data, sufficient training and ground truth data, system access, data access, and engagement. After examining these challenges in detail, the authors conclude there's a definitive "need for a more systemic change in how research in this domain is conducted, incentivized, supported, and regulated," with aspects such as stronger collaboration, more efficient machine learning methods, improved data curation, and improved data management methods.
In this 2023 article published in the journal F1000Research, Williamson et al. identify and discuss "eight key challenges in data management that must be addressed to further unlock the potential of [artificial intelligence] in crop and agronomic research." Noting the state of the agricultural research landscape and growing potentials of artificial intelligence and machine learning in the field, the authors perform a literature review that better shapes the nuances of those eight key challenges: data heterogeneity, data selection and digitization, data linkages, standardization and curation of data, sufficient training and ground truth data, system access, data access, and engagement. After examining these challenges in detail, the authors conclude there's a definitive "need for a more systemic change in how research in this domain is conducted, incentivized, supported, and regulated," with aspects such as stronger collaboration, more efficient machine learning methods, improved data curation, and improved data management methods.
 In this 2023 paper published in the journal High-Confidence Computing, Manisha and Jagadeeshwar present their custom food quality traceability system, which combines various tools like blockchain, internet of things (IoT) mechanisms, and deep learning architecture for improving traceability about perishable food supply chains (PFSCs). Noting previous works that incorporated some but not all of these elements, each with their own downsides, the authors chose a system model that incorporates blockchain-enabled RFID scans that, upon verification, get added to the overall secure ledger, along with associated IoT-base metadata from sensors gauging humidity and temperature. The duo turns to milk manufacturing and distribution in their case study, applied to their blockchain-enabled deep residual network (BC-DRN) methodology. After examining and comparing their results to other prevalent methodologies used in supply chain management (SCM). The authors conclude that their BC-DRN traceability system, when gauged on metrics like sensitivity, response time, and testing accuracy, beat out other methods. They add "the performance of the devised scheme can be improved by considering better feature extraction techniques. "
In this 2023 paper published in the journal High-Confidence Computing, Manisha and Jagadeeshwar present their custom food quality traceability system, which combines various tools like blockchain, internet of things (IoT) mechanisms, and deep learning architecture for improving traceability about perishable food supply chains (PFSCs). Noting previous works that incorporated some but not all of these elements, each with their own downsides, the authors chose a system model that incorporates blockchain-enabled RFID scans that, upon verification, get added to the overall secure ledger, along with associated IoT-base metadata from sensors gauging humidity and temperature. The duo turns to milk manufacturing and distribution in their case study, applied to their blockchain-enabled deep residual network (BC-DRN) methodology. After examining and comparing their results to other prevalent methodologies used in supply chain management (SCM). The authors conclude that their BC-DRN traceability system, when gauged on metrics like sensitivity, response time, and testing accuracy, beat out other methods. They add "the performance of the devised scheme can be improved by considering better feature extraction techniques. "
 In this brief 2023 article published in Journal of Clinical and Diagnostic Research, Patel et al. provide the results of their survey-based analysis of laboratorians' knowledge, attitude, and practice (KAP) towards laboratory quality through good clinical laboratory practice (GCLP) training. Noting the "ethical obligation to provide accurate and precise results that are cost- and time-effective," the authors state that it's imperative for clinical laboratory personnel to adhere to quality planning and system implementation, while also possessing an understanding of quality management principles as they apply to the lab. The authors describe their survey format and present the results of their survey, concluding that while no statistically significant differences could be found towards staff attitudes towards quality in the lab after GCLP training, laboratorians in their survey acknowledged the benefits of GCLP guidelines and accreditation, as well as the importance of training on such matters. "Such training and assessments would also aid in evaluating the performance of laboratory staff, contributing to improved learning, execution of GLPs, and consistent patient care services."
In this brief 2023 article published in Journal of Clinical and Diagnostic Research, Patel et al. provide the results of their survey-based analysis of laboratorians' knowledge, attitude, and practice (KAP) towards laboratory quality through good clinical laboratory practice (GCLP) training. Noting the "ethical obligation to provide accurate and precise results that are cost- and time-effective," the authors state that it's imperative for clinical laboratory personnel to adhere to quality planning and system implementation, while also possessing an understanding of quality management principles as they apply to the lab. The authors describe their survey format and present the results of their survey, concluding that while no statistically significant differences could be found towards staff attitudes towards quality in the lab after GCLP training, laboratorians in their survey acknowledged the benefits of GCLP guidelines and accreditation, as well as the importance of training on such matters. "Such training and assessments would also aid in evaluating the performance of laboratory staff, contributing to improved learning, execution of GLPs, and consistent patient care services."
 In this 2023 paper published in the journal Digital Discover, Scroggie et al. of the University of Sydney present their efforts towards utilizing the open developer platform GitHub as an electronic laboratory notebook (ELN) for chemistry research. Noting a lack of open-source ELNs with a focus on non-organic chemistry that have a wealth of collaboration tools, as well as problems with expensive and inflexible commercial ELNs, the authors turned to the many open facets of GitHub to repurpose its workings for synthetic chemistry projects. After a brief discussion of GitHub, the authors explain how they used the various facets of GitHub for ELN-related tasks, including notebooks, data and metadata management, and collaborative tools. They also acknowledged several shortcomings of their approach, including learning Markdown, dealing with data storage limitations, and integrating discipline-specific applications. The authors conclude that some of GitHub's features "are undeniably more oriented towards coders, such as the Actions tab in which users can set up workflows using code, these features do not detract from GitHub's usefulness as an ELN, which lies mainly in its adaptability and capacity for knowledge-sharing and collaboration."
In this 2023 paper published in the journal Digital Discover, Scroggie et al. of the University of Sydney present their efforts towards utilizing the open developer platform GitHub as an electronic laboratory notebook (ELN) for chemistry research. Noting a lack of open-source ELNs with a focus on non-organic chemistry that have a wealth of collaboration tools, as well as problems with expensive and inflexible commercial ELNs, the authors turned to the many open facets of GitHub to repurpose its workings for synthetic chemistry projects. After a brief discussion of GitHub, the authors explain how they used the various facets of GitHub for ELN-related tasks, including notebooks, data and metadata management, and collaborative tools. They also acknowledged several shortcomings of their approach, including learning Markdown, dealing with data storage limitations, and integrating discipline-specific applications. The authors conclude that some of GitHub's features "are undeniably more oriented towards coders, such as the Actions tab in which users can set up workflows using code, these features do not detract from GitHub's usefulness as an ELN, which lies mainly in its adaptability and capacity for knowledge-sharing and collaboration."
 In this 2023 paper published in the journal GigaScience, Nieminen et al. of the Berlin Institute of Health at Charité–Universitätsmedizin Berlin present SODAR (System for Omics Data Access and Retrieval), an open-source scientific data management system (SDMS) with a focus on omics data management during multiassay research studies. Noting numeroud data management challenges and dearth of open-source options, the authors describe the software framework, features, and limitations of the system. Highlighting SODAR's "programmable application programming interfaces (APIs) and command-line access for metadata and file storage," the authors conclude that SODAR can readily "support multiple technologies such as whole genome sequencing, single-cell sequencing, proteomics, and mass spectrometry," though some aspects such as automated data export and "data commons" access are currently not available.
In this 2023 paper published in the journal GigaScience, Nieminen et al. of the Berlin Institute of Health at Charité–Universitätsmedizin Berlin present SODAR (System for Omics Data Access and Retrieval), an open-source scientific data management system (SDMS) with a focus on omics data management during multiassay research studies. Noting numeroud data management challenges and dearth of open-source options, the authors describe the software framework, features, and limitations of the system. Highlighting SODAR's "programmable application programming interfaces (APIs) and command-line access for metadata and file storage," the authors conclude that SODAR can readily "support multiple technologies such as whole genome sequencing, single-cell sequencing, proteomics, and mass spectrometry," though some aspects such as automated data export and "data commons" access are currently not available.
 In this 2023 paper published in Healthcare Informatics Research, Chang et al. perform a literature review of current research in order to better characterize the currently viewed benefits of of artificial intelligence (AI), the internet of things (IoT), and personal health records (PHR) in the healthcare setting. Noting "limited empirical evidence regarding the benefits of information technology in healthcare settings," the authors examined 24 reviews and meta-analysis studies on these three technologies, with a strong focus on four outcome domains of clinical, psycho-behavioral, managerial, and socioeconomic implications. After detailing their findings, the authors conclude that "AI and PHRs can enhance clinical outcomes, while IoT holds promise for boosting managerial efficiency," though further research is required to address "the organizational and socioeconomic benefits of PHR," as well as the greater role of IoT in the healthcare setting.
In this 2023 paper published in Healthcare Informatics Research, Chang et al. perform a literature review of current research in order to better characterize the currently viewed benefits of of artificial intelligence (AI), the internet of things (IoT), and personal health records (PHR) in the healthcare setting. Noting "limited empirical evidence regarding the benefits of information technology in healthcare settings," the authors examined 24 reviews and meta-analysis studies on these three technologies, with a strong focus on four outcome domains of clinical, psycho-behavioral, managerial, and socioeconomic implications. After detailing their findings, the authors conclude that "AI and PHRs can enhance clinical outcomes, while IoT holds promise for boosting managerial efficiency," though further research is required to address "the organizational and socioeconomic benefits of PHR," as well as the greater role of IoT in the healthcare setting.
 Like many other parts of the food and beverage industry, the meat processing industry is guided by regulations that mandate the safety of the final product to the consumer. This includes laboratory testing for Trichinella spp., a nematode parasite responsible for the disease trichinosis. In order to ensure timely and accurate results, laboratories in the meat industry must act with purpose, implementing quality assurance (QA) practices that incorporate quality management systems (QMSs). In this 2023 journal article, Villegas-Pérez et al. examine the state of QA and QMSs in the in-house laboratories of Southern Spain's slaughterhouses and game-handling establishments, using canonical discriminant analysis (CDA) to gauge the effectiveness of those labs' practices. After lengthy review and discussion, the authors conclude that their CDA-based tools were able to discover "deficiencies in processes and procedures, necessitating measures for result reliability, due to facilities’ unfamiliarity with extensive QMS documentation." The authors offer several recommendations to fix those deficiencies.
Like many other parts of the food and beverage industry, the meat processing industry is guided by regulations that mandate the safety of the final product to the consumer. This includes laboratory testing for Trichinella spp., a nematode parasite responsible for the disease trichinosis. In order to ensure timely and accurate results, laboratories in the meat industry must act with purpose, implementing quality assurance (QA) practices that incorporate quality management systems (QMSs). In this 2023 journal article, Villegas-Pérez et al. examine the state of QA and QMSs in the in-house laboratories of Southern Spain's slaughterhouses and game-handling establishments, using canonical discriminant analysis (CDA) to gauge the effectiveness of those labs' practices. After lengthy review and discussion, the authors conclude that their CDA-based tools were able to discover "deficiencies in processes and procedures, necessitating measures for result reliability, due to facilities’ unfamiliarity with extensive QMS documentation." The authors offer several recommendations to fix those deficiencies.
 In this brief paper published in the EFSA Journal (from the European Food Safety Authority), Pineda-Pampliega et al. of the Norwegian Scientific Committee for Food and Environment (VKM) discuss the specifics of their work-related project to see "if existing commonly used [food safety] databases in risk assessment are in line with the FAIR principles" (which ensure data and information is findable, accessible, interoperable, and reusable), as well as to determine how to improve deficiencies in meeting FAIR principles. The authors describe their approach using data from both the EFSA's Chemical Hazards Database, OpenFoodTox, and the Institute of Marine Research's Seafood database. After evaluating these databases and describing the necessary framework activities to make them more FAIR, the authors conclude that their methods of using the R programming language, Shiny, GitHub, Zenodo, and appropriate file formats "is an essential step to ensuring the success of the future risk–benefit assessment [towards food safety], by offering more timely results with adequate spending of human and economic resources."
In this brief paper published in the EFSA Journal (from the European Food Safety Authority), Pineda-Pampliega et al. of the Norwegian Scientific Committee for Food and Environment (VKM) discuss the specifics of their work-related project to see "if existing commonly used [food safety] databases in risk assessment are in line with the FAIR principles" (which ensure data and information is findable, accessible, interoperable, and reusable), as well as to determine how to improve deficiencies in meeting FAIR principles. The authors describe their approach using data from both the EFSA's Chemical Hazards Database, OpenFoodTox, and the Institute of Marine Research's Seafood database. After evaluating these databases and describing the necessary framework activities to make them more FAIR, the authors conclude that their methods of using the R programming language, Shiny, GitHub, Zenodo, and appropriate file formats "is an essential step to ensuring the success of the future risk–benefit assessment [towards food safety], by offering more timely results with adequate spending of human and economic resources."
 In this 2023 paper published in JMIR Medical Informatics, Henke et al. of Technische Universität Dresden present the results of their effort to add "incremental loading" to the Medical Informatics in Research and Care in University Medicine's (MIRACUM's) clinical trial recruitment support systems (CTRSSs). Those CTRSSs already allows bulk loading of German-based FHIR data, supporting "the possibilities for multicentric and even international studies," but MIRACUM needed greater efficiencies when updating such data on a daily, incremental basis. The paper presents their literature review and approach to adding incremental loading to their systems. They conclude that the extract-transfer-load (ETL) "process no longer needs to be executed as a bulk load every day" with the change, instead being able to rely on "using bulk load for an initial load and switching to incremental load for daily updates." They add that this process has international applicability and is not limited to German FHIR data.
In this 2023 paper published in JMIR Medical Informatics, Henke et al. of Technische Universität Dresden present the results of their effort to add "incremental loading" to the Medical Informatics in Research and Care in University Medicine's (MIRACUM's) clinical trial recruitment support systems (CTRSSs). Those CTRSSs already allows bulk loading of German-based FHIR data, supporting "the possibilities for multicentric and even international studies," but MIRACUM needed greater efficiencies when updating such data on a daily, incremental basis. The paper presents their literature review and approach to adding incremental loading to their systems. They conclude that the extract-transfer-load (ETL) "process no longer needs to be executed as a bulk load every day" with the change, instead being able to rely on "using bulk load for an initial load and switching to incremental load for daily updates." They add that this process has international applicability and is not limited to German FHIR data.
 In this 2023 paper published in the Journal of Cannabis Research, Johnson et al. examine the current state of hemp-derived delta-9-tetrahydrocannabinol (Δ9-THC) products on the U.S. market after the passage of the Agriculture Improvement Act of 2018. Noting discrepancies and loopholes in the legislation, the authors performed laboratory analyses on 53 hemp-derived Δ9-THC products from 48 brands, while also examining aspects such as age verification, labeling consistency, and comparison of reported company values vs. analyzed values. After describing their methodology and results, the authors conclude that "the legal status of hemp-derived Δ9-THC products in America essentially permits their open sale while placing very few requirements on the companies selling them." The end result includes finding, for example, products "that have 3.7 times the THC content of edibles in adult-use states," as well as inaccurately labeled products.
In this 2023 paper published in the Journal of Cannabis Research, Johnson et al. examine the current state of hemp-derived delta-9-tetrahydrocannabinol (Δ9-THC) products on the U.S. market after the passage of the Agriculture Improvement Act of 2018. Noting discrepancies and loopholes in the legislation, the authors performed laboratory analyses on 53 hemp-derived Δ9-THC products from 48 brands, while also examining aspects such as age verification, labeling consistency, and comparison of reported company values vs. analyzed values. After describing their methodology and results, the authors conclude that "the legal status of hemp-derived Δ9-THC products in America essentially permits their open sale while placing very few requirements on the companies selling them." The end result includes finding, for example, products "that have 3.7 times the THC content of edibles in adult-use states," as well as inaccurately labeled products.
 "Artificial intelligence" (AI) may seem like a buzzword akin to the "nanotechnology" craze of the 2000s, but it is inevitably finding its way into scientific applications, including in the materials sciences. In this December 2022 article published in npj Computational Materials, Sbailò et al. present their AI-driven Novel Materials Discovery (NOMAD) toolkit as an extension of their NOMAD Repository & Archive, focused on making materials science data FAIR (findable, accessible, interoperable, and reusable), as well as AI-ready. After introducing the details of their workspace and goals towards adding notebook-based tools to the NOMAD Repository & Archive, the authors dive into the details of NOMAD AI Toolkit. They close by noting their toolkit "offers a selection of notebooks demonstrating such [AI-based] workflows, so that users can understand step by step what was done in publications and readily modify and adapt the workflows to their own needs." They add that the system "will allow for enhanced reproducibility of data-driven materials science papers and dampen the learning curve for newcomers to the field."
"Artificial intelligence" (AI) may seem like a buzzword akin to the "nanotechnology" craze of the 2000s, but it is inevitably finding its way into scientific applications, including in the materials sciences. In this December 2022 article published in npj Computational Materials, Sbailò et al. present their AI-driven Novel Materials Discovery (NOMAD) toolkit as an extension of their NOMAD Repository & Archive, focused on making materials science data FAIR (findable, accessible, interoperable, and reusable), as well as AI-ready. After introducing the details of their workspace and goals towards adding notebook-based tools to the NOMAD Repository & Archive, the authors dive into the details of NOMAD AI Toolkit. They close by noting their toolkit "offers a selection of notebooks demonstrating such [AI-based] workflows, so that users can understand step by step what was done in publications and readily modify and adapt the workflows to their own needs." They add that the system "will allow for enhanced reproducibility of data-driven materials science papers and dampen the learning curve for newcomers to the field."
 In this 2023 article published in Journal of Clinical and Diagnostic Research, Naphade et al. provide a brief introductory-level review of the importance of quality control (QC) to the clinical laboratory. After a brief introduction on clinical lab testing, the authors analyze the wide variety of sources for laboratory errors, covering the pre-analytical, analytical. and post-analytical phases. They then introduce the concept of quality control, followed by explaining how QC is implemented in the laboratory, including through the use of QC materials, statistical control charts, and shifts and trends. They conclude this review by stating that "reliable and confident laboratory testing avoids misdiagnosis, delayed treatment, and unnecessary costing of repeat testing." They add that given these benefits, "the individual laboratory should assess and analyze their own QC process to find out the possible root cause of any digressive test results which are not correlating with patients' clinical presentation or expected response to treatment."
In this 2023 article published in Journal of Clinical and Diagnostic Research, Naphade et al. provide a brief introductory-level review of the importance of quality control (QC) to the clinical laboratory. After a brief introduction on clinical lab testing, the authors analyze the wide variety of sources for laboratory errors, covering the pre-analytical, analytical. and post-analytical phases. They then introduce the concept of quality control, followed by explaining how QC is implemented in the laboratory, including through the use of QC materials, statistical control charts, and shifts and trends. They conclude this review by stating that "reliable and confident laboratory testing avoids misdiagnosis, delayed treatment, and unnecessary costing of repeat testing." They add that given these benefits, "the individual laboratory should assess and analyze their own QC process to find out the possible root cause of any digressive test results which are not correlating with patients' clinical presentation or expected response to treatment."
 In this 2023 paper published in the journal Scientific Data, Ghiringhelli et al. present the results of an international workshop of materials scientists who came together to discuss metadata and data formats as they relate to data sharing in materials science. With a focus on the FAIR principles (findable, accessible, interoperable, and reusable), the authors introduce the concept of materials science data management needs, and the value of metadata towards making shareable data FAIR. They then go into specific use cases, such as electronic-structure calculations, potential-energy sampling, and managing metadata for computational workflows. They also address the nuances of file formats and metadata schemas as they relate to those file formats. They clise with an outlook on the use of ontologies in materials science, followed by conclusions, with a heavy emphasis on "the importance of developing a hierarchical and modular metadata schema in order to represent the complexity of materials science data and allow for access, reproduction, and repurposing of data, from single-structure calculations to complex workflows." They add: "the biggest benefit of meeting the interoperability challenge will be to allow for routine comparisons between computational evaluations and experimental observations."
In this 2023 paper published in the journal Scientific Data, Ghiringhelli et al. present the results of an international workshop of materials scientists who came together to discuss metadata and data formats as they relate to data sharing in materials science. With a focus on the FAIR principles (findable, accessible, interoperable, and reusable), the authors introduce the concept of materials science data management needs, and the value of metadata towards making shareable data FAIR. They then go into specific use cases, such as electronic-structure calculations, potential-energy sampling, and managing metadata for computational workflows. They also address the nuances of file formats and metadata schemas as they relate to those file formats. They clise with an outlook on the use of ontologies in materials science, followed by conclusions, with a heavy emphasis on "the importance of developing a hierarchical and modular metadata schema in order to represent the complexity of materials science data and allow for access, reproduction, and repurposing of data, from single-structure calculations to complex workflows." They add: "the biggest benefit of meeting the interoperability challenge will be to allow for routine comparisons between computational evaluations and experimental observations."
 In this 2023 journal article published in International Journal of Molecular Sciences, Jadhav et al. recommend a more comprehensive approach to analyzing the Cannabis plant, its extracts, and its constituents, turning to the still-evolving concept of authentomics—"which involves combining the non-targeted analysis of new samples with big data comparisons to authenticated historic datasets"—of the food industry as a paradigm worth shifting towards. After an introduction on metabolomics and a review of its various technologies, the authors examine the current state of cannabis laboratory testing, how its regulated and standardized, and a variety of issues that make cannabis testing less consistent, as well as how cannabis can become adulterated in the grow chain. They then discuss authentomics and apply it to the cannabis industry, as well address what considerations need to be made in the future to make the most of authentomics. The author conclude that an authentomics approach "provides a robust method for verifying the quality of cannabis products."
In this 2023 journal article published in International Journal of Molecular Sciences, Jadhav et al. recommend a more comprehensive approach to analyzing the Cannabis plant, its extracts, and its constituents, turning to the still-evolving concept of authentomics—"which involves combining the non-targeted analysis of new samples with big data comparisons to authenticated historic datasets"—of the food industry as a paradigm worth shifting towards. After an introduction on metabolomics and a review of its various technologies, the authors examine the current state of cannabis laboratory testing, how its regulated and standardized, and a variety of issues that make cannabis testing less consistent, as well as how cannabis can become adulterated in the grow chain. They then discuss authentomics and apply it to the cannabis industry, as well address what considerations need to be made in the future to make the most of authentomics. The author conclude that an authentomics approach "provides a robust method for verifying the quality of cannabis products."
 In this 2023 article published in Science and Technology of Advanced Materials: Methods, Ishii et al. describe their cross-institutional efforts towards integrating X-ray absorption fine structure (XAFS) spectra and associated metadata across multiple databases to improve materials informatics processes and research methods of materials discovery and analysis. Their new public database, MDR XAFS DB, was developed towards this goal, while addressing two main issues with unifying such data: the difficulties of "designing and collecting metadata describing spectra and sample details, and unifying the vocabulary used in the metadata, including not only metadata items (keys) but also descriptions (values)." After discussing the system's construction and contents, the authors conclude their system "has achieved seamless cross searchability with the use of sample nomenclature so that database users do not have to be aware of the differences in the local metadata of the facilities that provide the data." Though still with some challenges to address, the authors add that while the culture of open data hasn't truly taken hold in materials science, they hope "that this initiative will be a trigger to promote the utilization of materials data."
In this 2023 article published in Science and Technology of Advanced Materials: Methods, Ishii et al. describe their cross-institutional efforts towards integrating X-ray absorption fine structure (XAFS) spectra and associated metadata across multiple databases to improve materials informatics processes and research methods of materials discovery and analysis. Their new public database, MDR XAFS DB, was developed towards this goal, while addressing two main issues with unifying such data: the difficulties of "designing and collecting metadata describing spectra and sample details, and unifying the vocabulary used in the metadata, including not only metadata items (keys) but also descriptions (values)." After discussing the system's construction and contents, the authors conclude their system "has achieved seamless cross searchability with the use of sample nomenclature so that database users do not have to be aware of the differences in the local metadata of the facilities that provide the data." Though still with some challenges to address, the authors add that while the culture of open data hasn't truly taken hold in materials science, they hope "that this initiative will be a trigger to promote the utilization of materials data."
 Forensic laboratories are largely bound to performing their actions with quality highly in mind. In fact, standards like ISO/IEC 17025 help such labs focus more on quality with the help of a quality management system (QMS). But even with a strong focus on quality, inconsistencies in recording, managing, and reporting quality issues can be problematic. To that point, Heavey et al. examine government-based forensic service provider agencies in Australia and New Zealand in this 2023 journal article, using the survey format to gain insight into potential QMS weaknesses. After thoroughly reviewing the methodology and results for their survey, the authors conclude that "the need for further research into the standardization of systems underpinning the management of quality issues in forensic science" is evident from their research, with a greater need "to support transparent and reliable justice outcomes" through improving "evidence-based standard terminology for quality issues in forensic science to support data sharing and reporting, enhance understanding of quality issues, and promote transparency in forensic science."
Forensic laboratories are largely bound to performing their actions with quality highly in mind. In fact, standards like ISO/IEC 17025 help such labs focus more on quality with the help of a quality management system (QMS). But even with a strong focus on quality, inconsistencies in recording, managing, and reporting quality issues can be problematic. To that point, Heavey et al. examine government-based forensic service provider agencies in Australia and New Zealand in this 2023 journal article, using the survey format to gain insight into potential QMS weaknesses. After thoroughly reviewing the methodology and results for their survey, the authors conclude that "the need for further research into the standardization of systems underpinning the management of quality issues in forensic science" is evident from their research, with a greater need "to support transparent and reliable justice outcomes" through improving "evidence-based standard terminology for quality issues in forensic science to support data sharing and reporting, enhance understanding of quality issues, and promote transparency in forensic science."
 In this brief 2023 article published in the journal Tomography, Michele Larobina of Consiglio Nazionale delle Ricerche provides background on "the innovation, advantages, and limitations of adopting DICOM and its possible future directions." After an introduction on the Digital Imaging and Communications in Medicine standard, Larobina then emphasizes the benefits of the standard's approach to handling image metadata, followed by the exchange component of the standard. Larobina then examines the strengths and weaknesses of the DICOM standard, before concluding that DICOM has not only demonstrated "the positive influence and added value that a standard can have in a specific field," but also "encouraged and facilitated data exchange between researchers, creating added value for research." Despite this, "the time is ripe to review some of the initial directives" of DICOM, says Larobina, in order for the the standard to remain relevant going forward.
In this brief 2023 article published in the journal Tomography, Michele Larobina of Consiglio Nazionale delle Ricerche provides background on "the innovation, advantages, and limitations of adopting DICOM and its possible future directions." After an introduction on the Digital Imaging and Communications in Medicine standard, Larobina then emphasizes the benefits of the standard's approach to handling image metadata, followed by the exchange component of the standard. Larobina then examines the strengths and weaknesses of the DICOM standard, before concluding that DICOM has not only demonstrated "the positive influence and added value that a standard can have in a specific field," but also "encouraged and facilitated data exchange between researchers, creating added value for research." Despite this, "the time is ripe to review some of the initial directives" of DICOM, says Larobina, in order for the the standard to remain relevant going forward.
 In this article published in BMC Medical Education, Xu et al. present their four-stage clinical biochemistry teaching methodology which incorporates elements of the International Organization for Standardization's ISO 15189 standard. Noting a lack of student awareness of ISO 15189 Medical laboratories — Requirements for quality and competence and the value its emphasis on quality management brings to laboratories, the authors set upon incorporating the standard into a case-based learning (CBL) approach. After the authors describe the details of their approach and the survey-based results analysis of its outcomes, the authors conclude that outside of improving student outcomes in clinical biochemistry, "ISO 15189 concepts of continuous improvement were implanted, thereby putting the concept of plan–do–check–act (PDCA) into practice, developing recording habits, and improving communication skills."
In this article published in BMC Medical Education, Xu et al. present their four-stage clinical biochemistry teaching methodology which incorporates elements of the International Organization for Standardization's ISO 15189 standard. Noting a lack of student awareness of ISO 15189 Medical laboratories — Requirements for quality and competence and the value its emphasis on quality management brings to laboratories, the authors set upon incorporating the standard into a case-based learning (CBL) approach. After the authors describe the details of their approach and the survey-based results analysis of its outcomes, the authors conclude that outside of improving student outcomes in clinical biochemistry, "ISO 15189 concepts of continuous improvement were implanted, thereby putting the concept of plan–do–check–act (PDCA) into practice, developing recording habits, and improving communication skills."
 In this 2023 article published in the journal Water, Marebesi et al. of the University of Georgia present their findings on phytoremediation of Cadmium (Cd)-contaminated soils using day-neutral and photoperiod-sensitive hemp plants. After a thorough introduction on the topic of using Cannabis plants for phytoremediation (including the flowers), the authors present their materials and methods, focusing on Apricot Auto, Alpha Explorer, Von, and T1 varieties of hemp. They then present the results of their experiments, addressing plant height, biomass yield, Cd concentrations in various plant tissues, nutrient partitioning, and THC/CBD concentrations. They conclude noting that "while Cd concentration was significantly higher in roots, all four varieties were efficient in translocating Cd from roots to shoots." They also conclude that THC/CBD loss is affected by remediation in some cases, as was height and biomass. Finally, they highlight the importance of Cd testing of flowers for medical markets where the plant is used.
In this 2023 article published in the journal Water, Marebesi et al. of the University of Georgia present their findings on phytoremediation of Cadmium (Cd)-contaminated soils using day-neutral and photoperiod-sensitive hemp plants. After a thorough introduction on the topic of using Cannabis plants for phytoremediation (including the flowers), the authors present their materials and methods, focusing on Apricot Auto, Alpha Explorer, Von, and T1 varieties of hemp. They then present the results of their experiments, addressing plant height, biomass yield, Cd concentrations in various plant tissues, nutrient partitioning, and THC/CBD concentrations. They conclude noting that "while Cd concentration was significantly higher in roots, all four varieties were efficient in translocating Cd from roots to shoots." They also conclude that THC/CBD loss is affected by remediation in some cases, as was height and biomass. Finally, they highlight the importance of Cd testing of flowers for medical markets where the plant is used.
 As technological, cultural, and environmental shifts continue to put pressure on the ways academic and private research is conducted, researchers must further strive to get the most out of their work. This holds true across a broad swath of scientific fields, including materials science. Noting the continuing expansion of laboratory automation and software-based tools like artificial intelligence (AI) and machine learning (ML), Tamura et al. emphasize that materials science and novel materials research can benefit greatly from adopting these tools. In this 2023 paper, the authors present their take on this trend, in the form of NIMS-OS, a Python-based system developed to take advantage of completely automated material discovery experiments. After presenting an introduction on the topic, the authors discuss the development philosophy and approach to making NIMS-OS, while also describing use of both the Python command-line tool and the graphical user interface (GUI)-driven version of the software. The authors conclude that their system demonstrates how "various problems for automated materials exploration to be commonly performed," while at the same time being "designed to be easily adaptable to a variety of original robotic systems in materials science."
As technological, cultural, and environmental shifts continue to put pressure on the ways academic and private research is conducted, researchers must further strive to get the most out of their work. This holds true across a broad swath of scientific fields, including materials science. Noting the continuing expansion of laboratory automation and software-based tools like artificial intelligence (AI) and machine learning (ML), Tamura et al. emphasize that materials science and novel materials research can benefit greatly from adopting these tools. In this 2023 paper, the authors present their take on this trend, in the form of NIMS-OS, a Python-based system developed to take advantage of completely automated material discovery experiments. After presenting an introduction on the topic, the authors discuss the development philosophy and approach to making NIMS-OS, while also describing use of both the Python command-line tool and the graphical user interface (GUI)-driven version of the software. The authors conclude that their system demonstrates how "various problems for automated materials exploration to be commonly performed," while at the same time being "designed to be easily adaptable to a variety of original robotic systems in materials science."







