Posted on April 25, 2023
By LabLynx
Journal articles

In this 2023 article published in the journal
Processes, Frede
et al. of TU Dortmund University work with analytical consultancy d-fine GmbH to demonstrate experimental workflows of a microscale reaction calorimeter over d-fine's open-source internet of things (IoT) software platform. Their goal: provide an example of how data collection and capture methods can and should be standardized across an automated laboratory. After a brief introduction, the authors present their materials and methods, along with a case study of hydrolysis of acetic anhydride. After presenting the results of their case study, they conclude that "[t]he modular data platform complements the reaction calorimeter very well with additional functionalities," and by adding even more instruments, "[t]his approach will further harmonize data management within our laboratory while better applying FAIR Guiding Principles."
Posted on April 18, 2023
By LabLynx
Journal articles
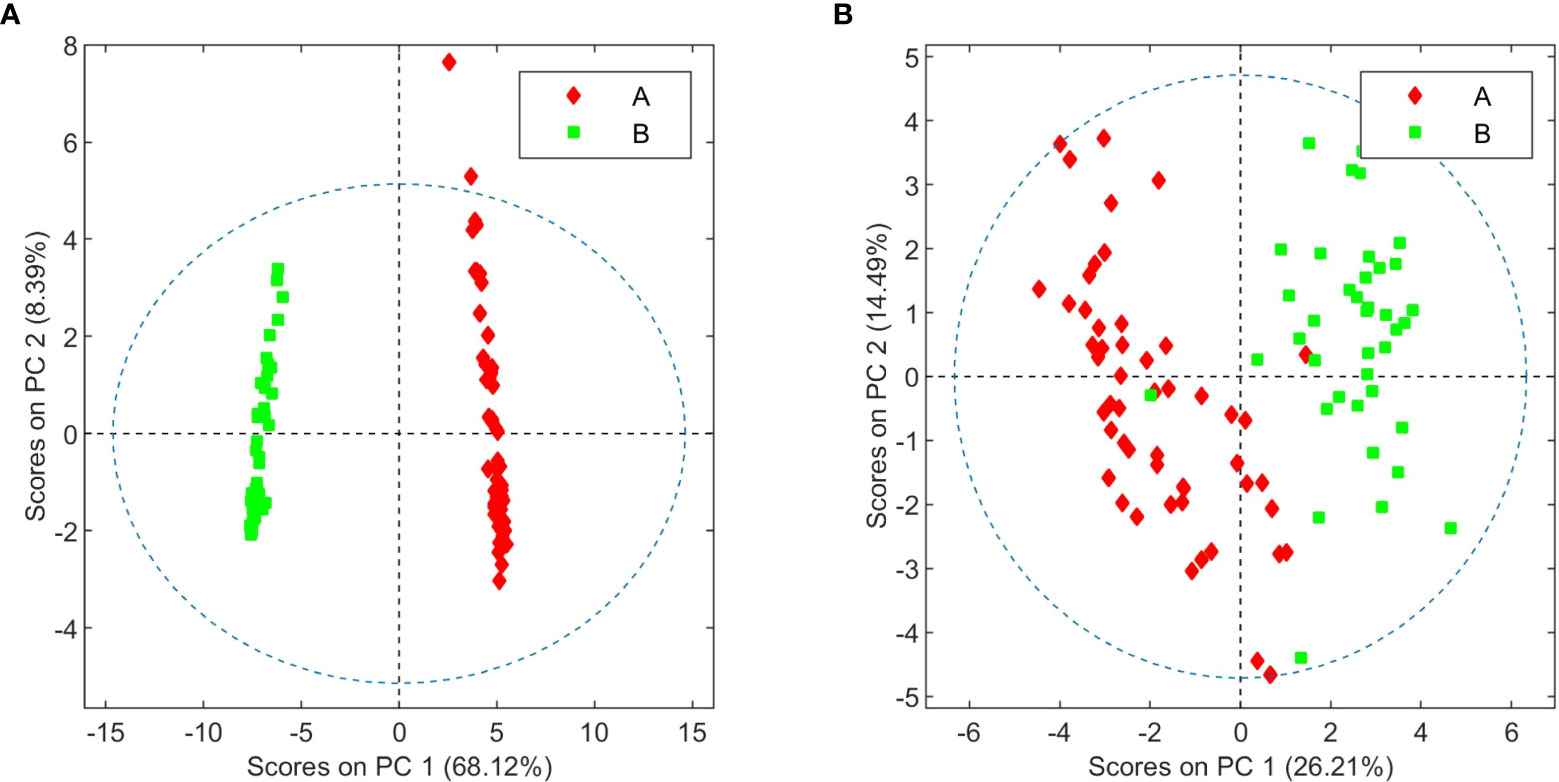
In this 2023 paper published in the journal
Frontiers in Plant Science, Fernández
et al. propose that
Cannabis should be characterized "according to its chemical composition (i.e., its metabolome) and not only its botanical traits," while conducting a series of experiments to back their claim. After an introduction to cannabis and its chemical characterization, the authors describe the materials, methods, and results of their experimentation using metabolomics on high-THCA and high-CBDA
C. sativa chemovars with medicinal potential. They conclude that many "differences between the two varieties beyond their cannabinoid composition" could be found through this experimentation, and that these differences could be "instrumental to breeders in the development of plant varieties agronomically adapted to specific environmental conditions." They also noted that fungal infections changed the metabolome of tested plants, which can better aid in the "classification of healthy and diseased plants."
Posted on April 11, 2023
By LabLynx
Journal articles
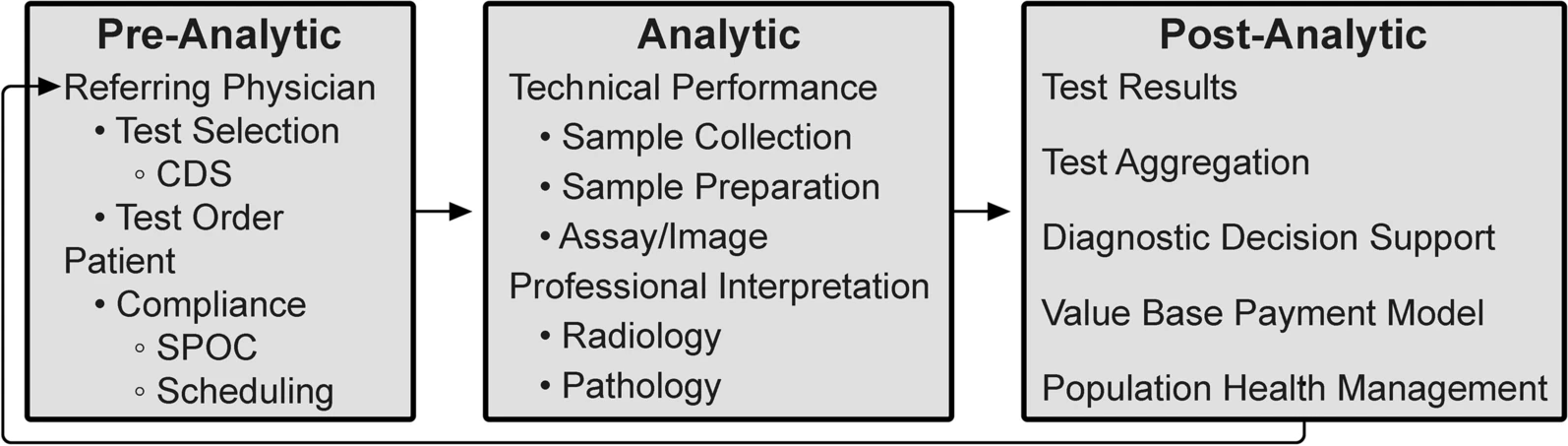
In this 2022 paper published in
Insights into Imaging, Beauchamp
et al. focus in on the promise of "integrative diagnostics" as a means towards aggregating diagnostic data for more meaningful interpretation and contextualization, by extension translating to directly relatable clinical action. Looking specifically at radiology and pathology workflows, the authors describe what integrative diagnostics looks like, how the process should work, and what promise it holds for
in vivo and
in vitro radiology and pathology workflows. The authors turn to 24 different sources in highlighting the clinical potential of integrative diagnostics for a wide variety of diseases, highlighting the added clinical value and improved discovery possible with integrative diagnostics' application. After examining the current state of the art, the authors conclude by making several recommendations towards implementing integrative diagnostics, while also noting that "[c]reating financial models that demonstrate the economic value proposition of [integrative diagnostics] will be a necessary catalyst for change" for the overall health care system.
Posted on April 3, 2023
By LabLynx
Journal articles
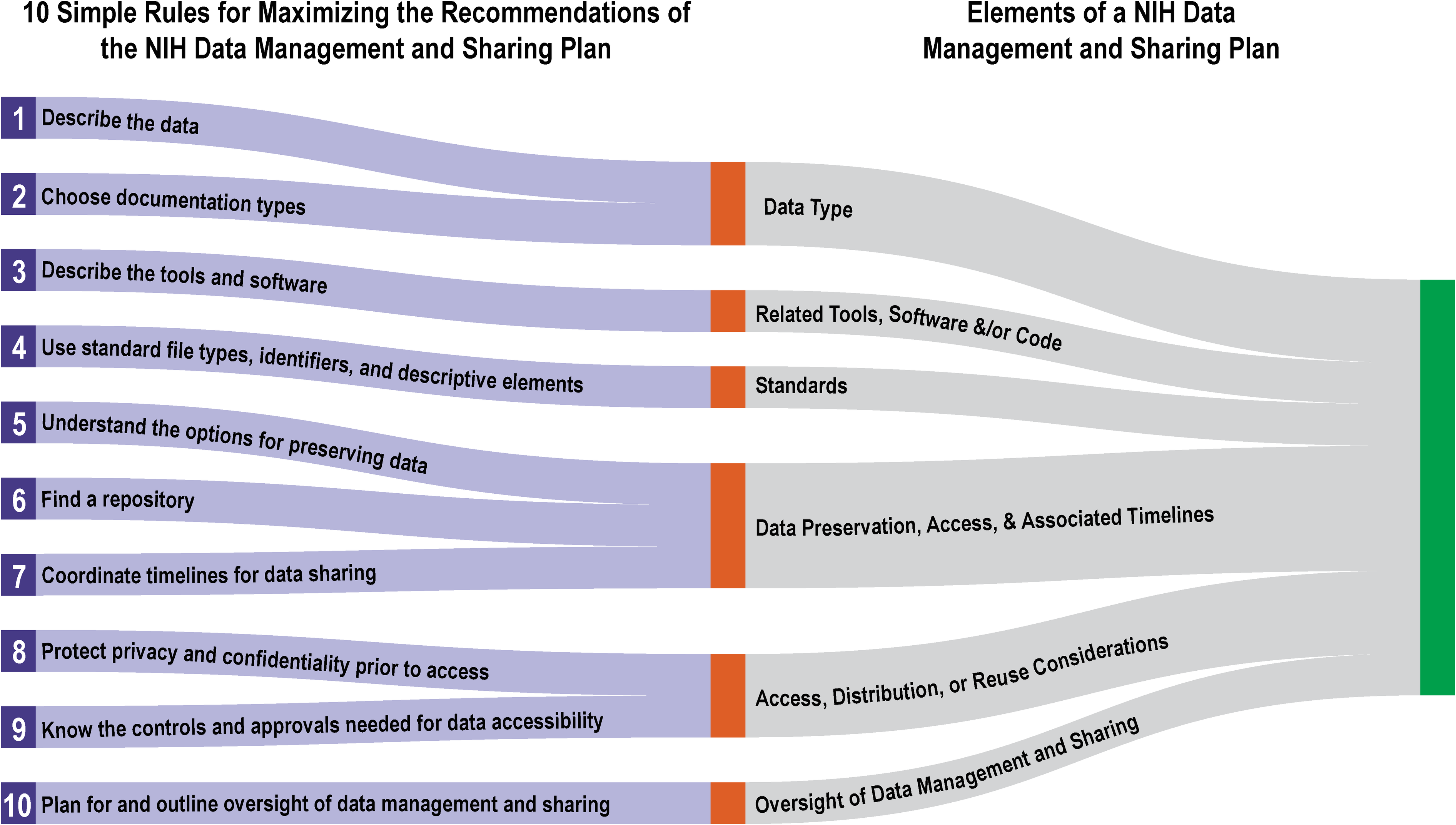
In January 2023, the National Institutes of Health's (NIH's) Policy for Data Management and Sharing came into effect, requiring guidance on elements "for the submission of a data management and sharing plan (DMSP)" for NIH-funded projects. While the NIH created supplementary guidance to developing a DMSP, some ambiguity arguably remained. In this 2022 article published in
PLOS Computational Biology, Gonzales
et al. provide additional insight into the process. After a brief introduction, the authors present 10 helpful tips towards getting the most of a DMSP in complying with the NIH and its requirements. They conclude by examining the various stakeholders involved with the DMSP process and finding that "[g]ood DMSP practices can support better engagement with and accountability to the public who benefit from research."
Posted on March 27, 2023
By LabLynx
Journal articles
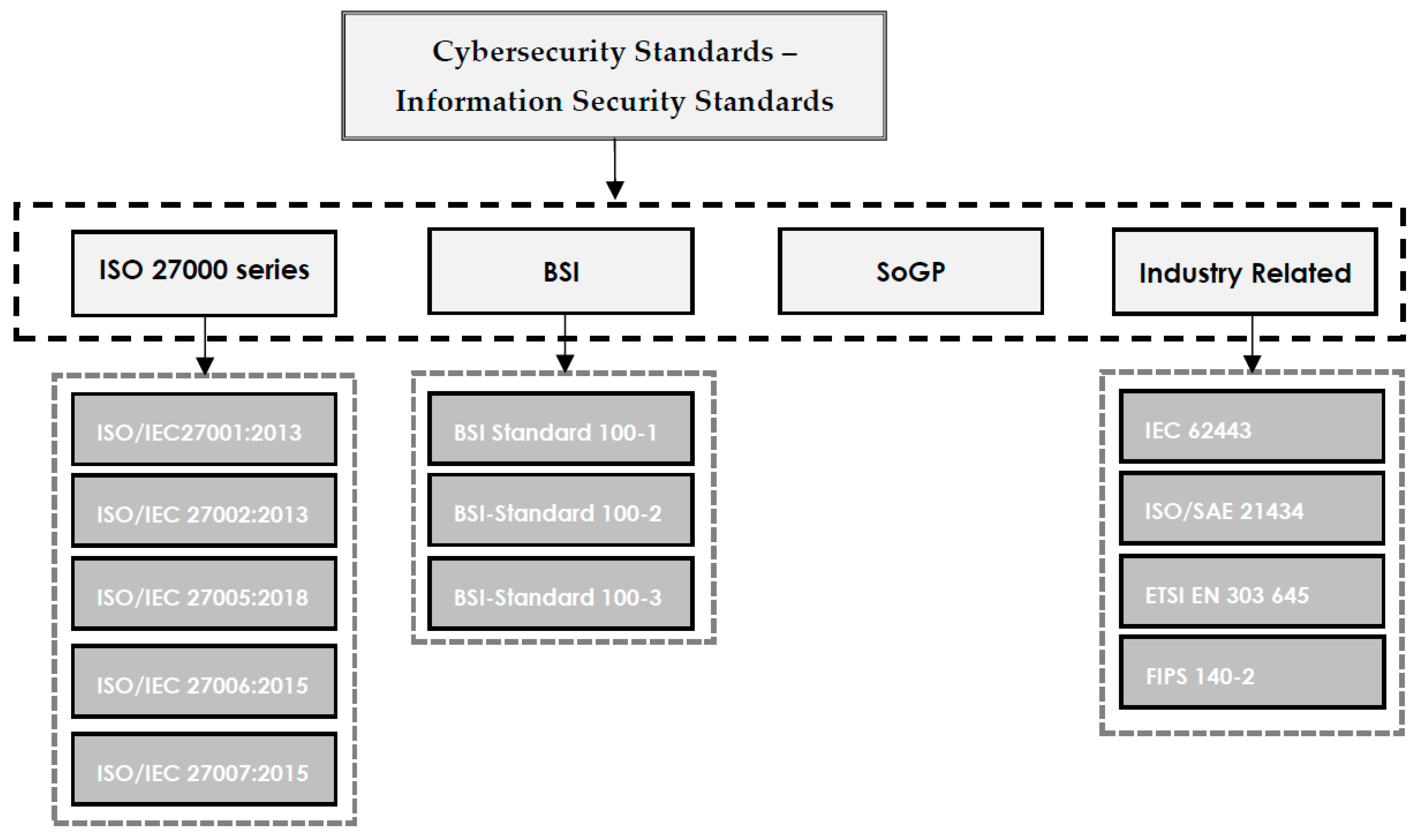
In this 2022 journal article published in
Electronics, University Canada West's Hamed Taherdoost presents a narrative literature review of a variety of cybersecurity standards and frameworks as a means to "help organizations select the cybersecurity standard or framework that best fits their cybersecurity requirements." After providing a brief introduction, Taherdoost speaks generally about cybersecurity standards and frameworks, and what they help organizations of all types achieve with their own cybersecurity. The author then goes into greater detail of those standards and frameworks, as well as their supporting documentation. Taherdoost then presents the methodology for his literature review, followed by a discussion of the results of that review. The author concludes that the presented standards and frameworks help organizations "ensure the security of data against cyber threats," though one standard or framework "may not fulfill all the demands of an organization, and it may be necessary to employ a combination of standards in order to ensure security against cyber threats and data loss."
Posted on March 20, 2023
By LabLynx
Journal articles
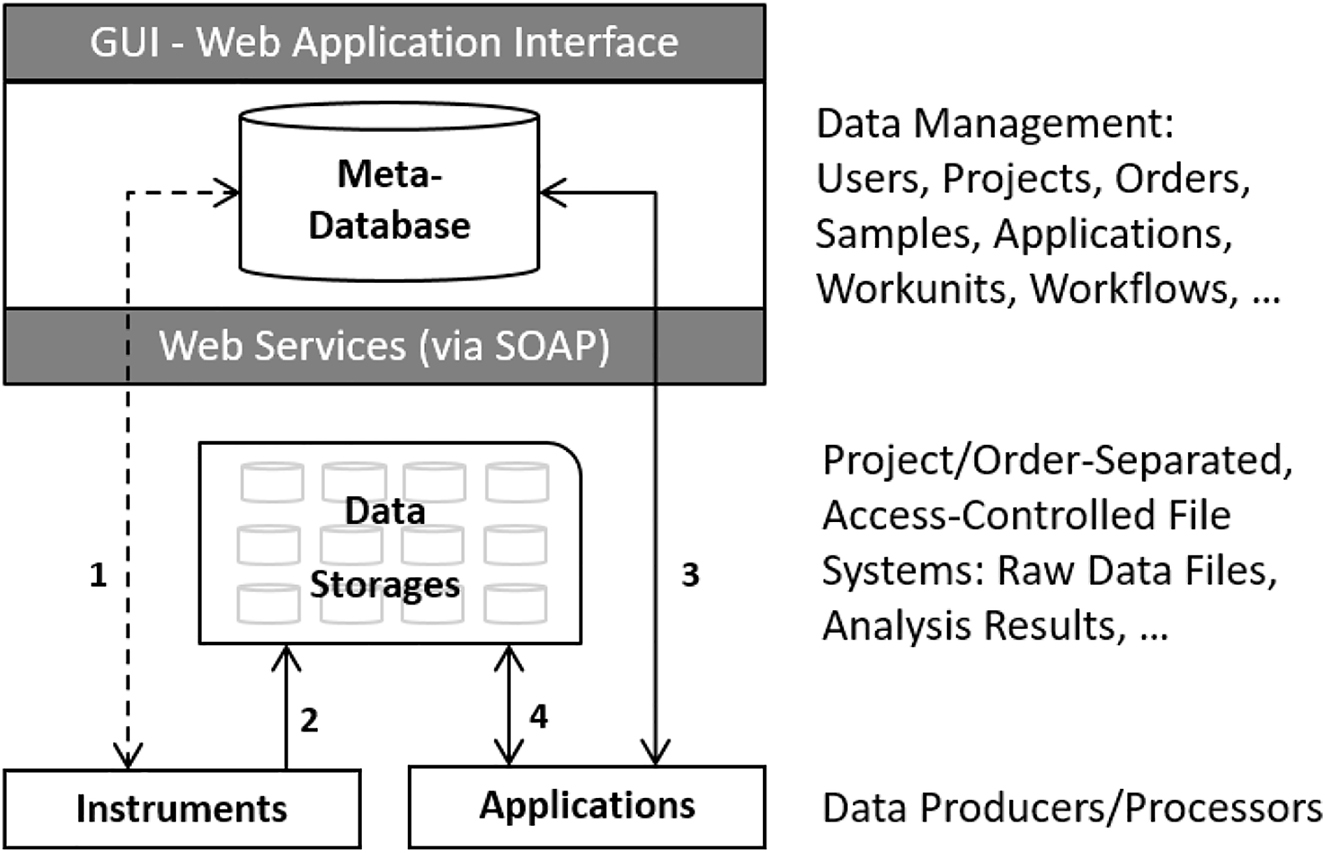
In this 2022 journal article published in
Journal of Integrative Bioinformatics, Panse
et al. discuss the state of research-based core facilities that share research resources across institutions or organizations, in particular in regards to data and information management. The authors note that the "interdisciplinary nature of such scientific projects requires a powerful and flexible platform for data annotation, communication, and exploration," and the authors—associated with the Functional Genomics Center Zurich (FGCZ)—present B-Fabric, their solution to data and information management in such settings. After providing a brief background, the authors discuss the development and application of B-Fabric at FGCZ, along with lessons learned. They conclude that with systems like B-Fabric, "ad-hoc visualizations and analytics can be realized on top of an integrative platform that captures and processes any kind of data and scientific processes." The platform is also able "to decouple metadata management from data processing and visualization components" while also optimizing specific software software environments for different research areas.
Posted on March 13, 2023
By LabLynx
Journal articles
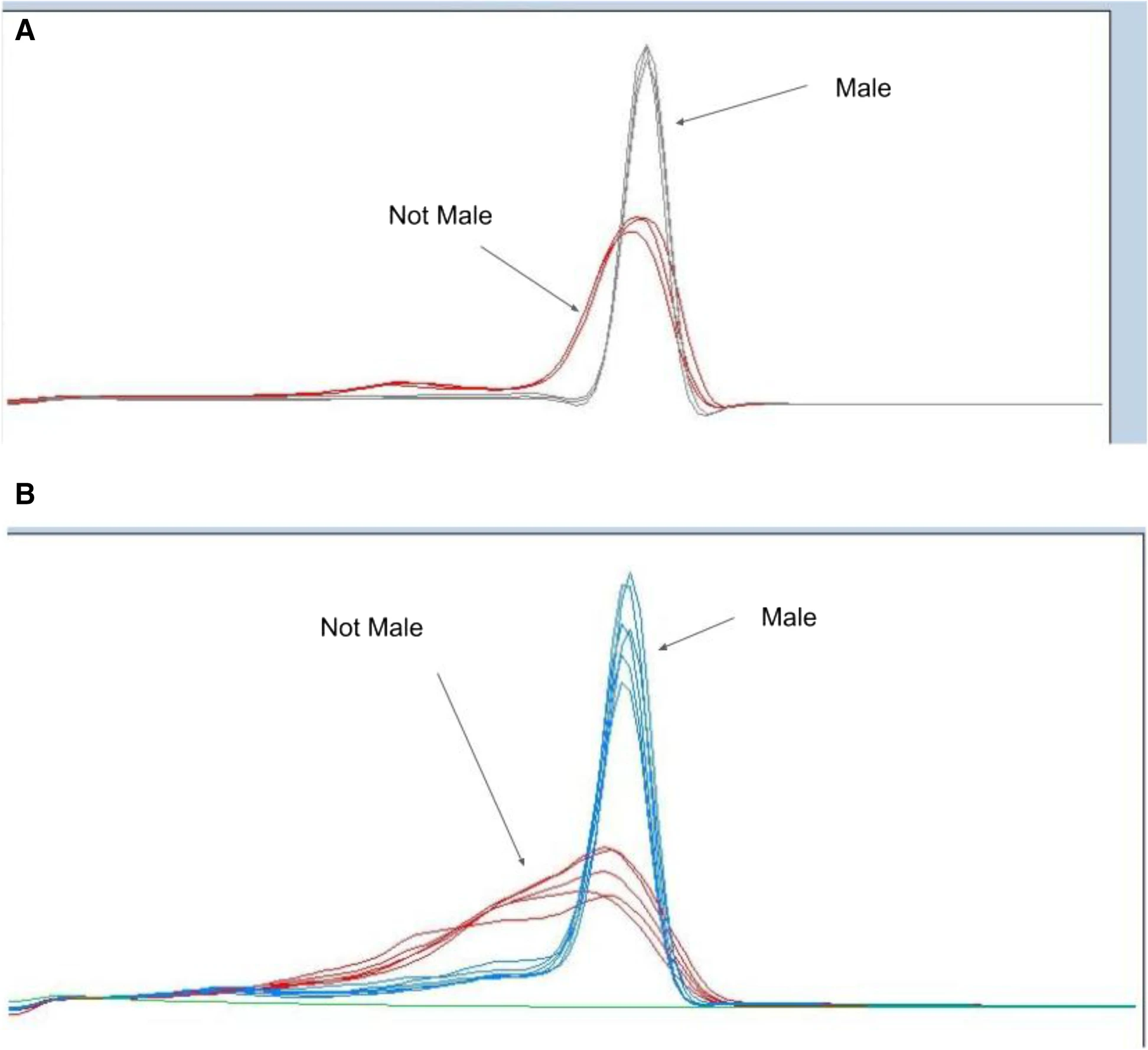
Genotyping of
Cannabis sativa is a useful component of not only gender identification of plants for cultivators but also other types of laboratory testing and research of
Cannabis. Performing such tests in a reliable, cost-effective, and high-performance way has significant value for
Cannabis testing laboratories. In this 2022 journal article, Torres
et al. compare a number of different genotyping techniques, ultimately demonstrating some of that value. After providing background information and their methodology, the authors discuss the results of gender testing using a wide array of genotyping techniques. They conclude that "the multiple methodologies presented here allow for accurate, quick, and cost-effective screening that will enable future development of germplasm and the industry." They add that their "real-time assays can be performed by cannabis testing laboratories performing diagnostic testing, as well as in the field with minimal molecular biology equipment and expertise."
Posted on March 6, 2023
By LabLynx
Journal articles
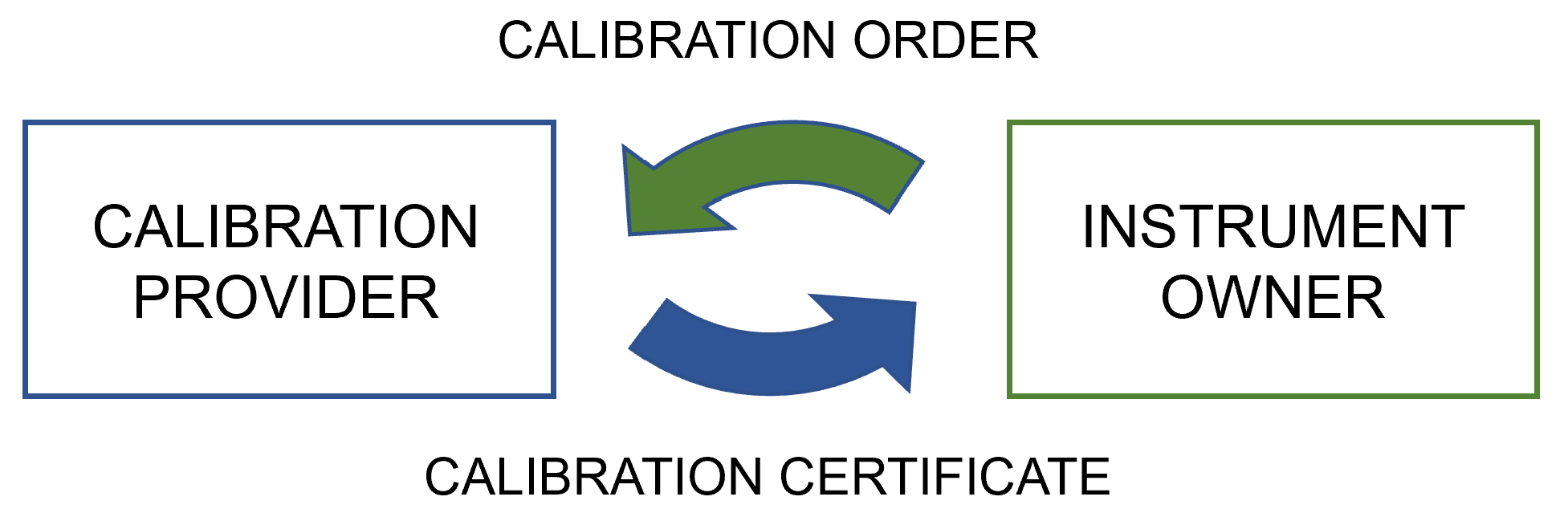
In this 2022 journal article published in
Applied Sciences, Mustapää
et al. turn to the calibration process of the pharmaceutical industry—as well as their regulatory- and standards-based requirements—as inspiration for building a proof-of-concept software platform that better ensures "the preservation of traceability and data integrity" of calibration data. After a lengthy and thorough background to the topic of laboratory calibrations and calibration data management, the authors present their proof of concept software platform, designed "to harmonize the data exchange between the parties of a calibration chain to enable automation of the data management processes in the receiving end." They then discuss the results of their efforts, challenges in advancing such metrology infrastructure, and topics that require further research. They conclude that the real use cases tested on their proof of concept "proved that the multitenant platform is an efficient means of organizing the exchange of [digital calibration certificates]," and that "the system would suit the requirements of other industries as well, providing cost-efficiency through economies of scale."
Posted on February 27, 2023
By LabLynx
Journal articles
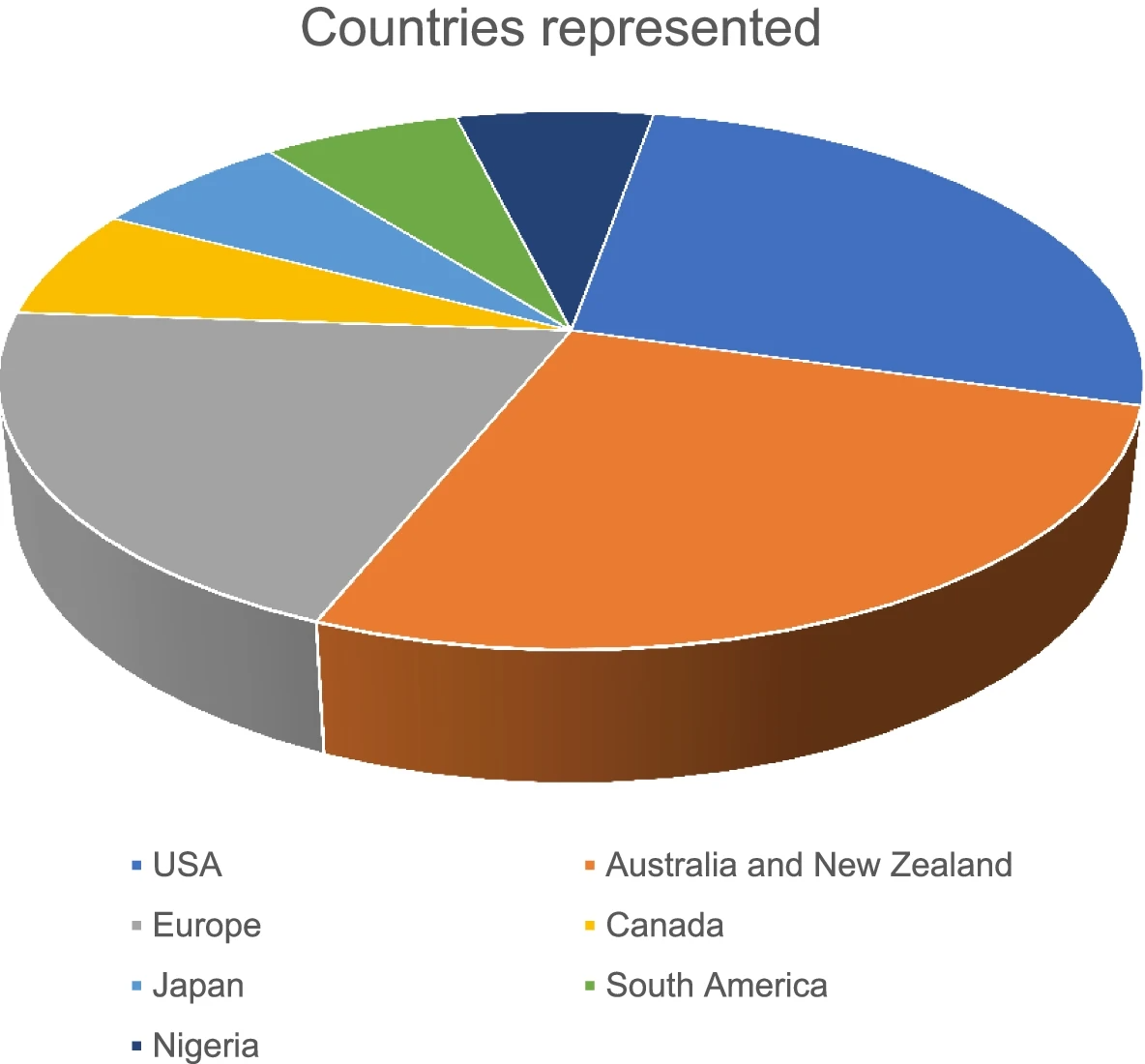
In this 2022 paper published in the
Egyptian Journal of Forensic Sciences, Prahladh and van Wyk provide a first-of-its-kind comprehensive scoping review of "articles involving unnatural deaths, focused on data practice or data management systems, relating to forensic medicine, all study designs, and published in English," with the goal of being able to draw conclusions about the current academic view of forensic data management. After examining more than 23,000 articles, the authors were only able to find 16 that met their criteria, according to their stated methodology. After a thorough review of those 16 articles, the duo concludes that "the literature shows that electronic data reporting systems are relevant and were developed from the recognition of coronial data as not only a part of the death investigation but as a contributor to preventable death research and public health initiatives." However, there are few lower- and middle-income countries (LMICs) represented in the study, and the authors believe that given how "forensic medicine departments can utilize simple and available tools that can advance standardization of data collection, storage, and reporting," it would behoove more LMICs to consider pilot systems such as "the Nigerian pilot program [that] still managed to contribute to statistically relevant data for impactful research" while using low-cost and readily available data management tools.
Posted on February 21, 2023
By LabLynx
Journal articles
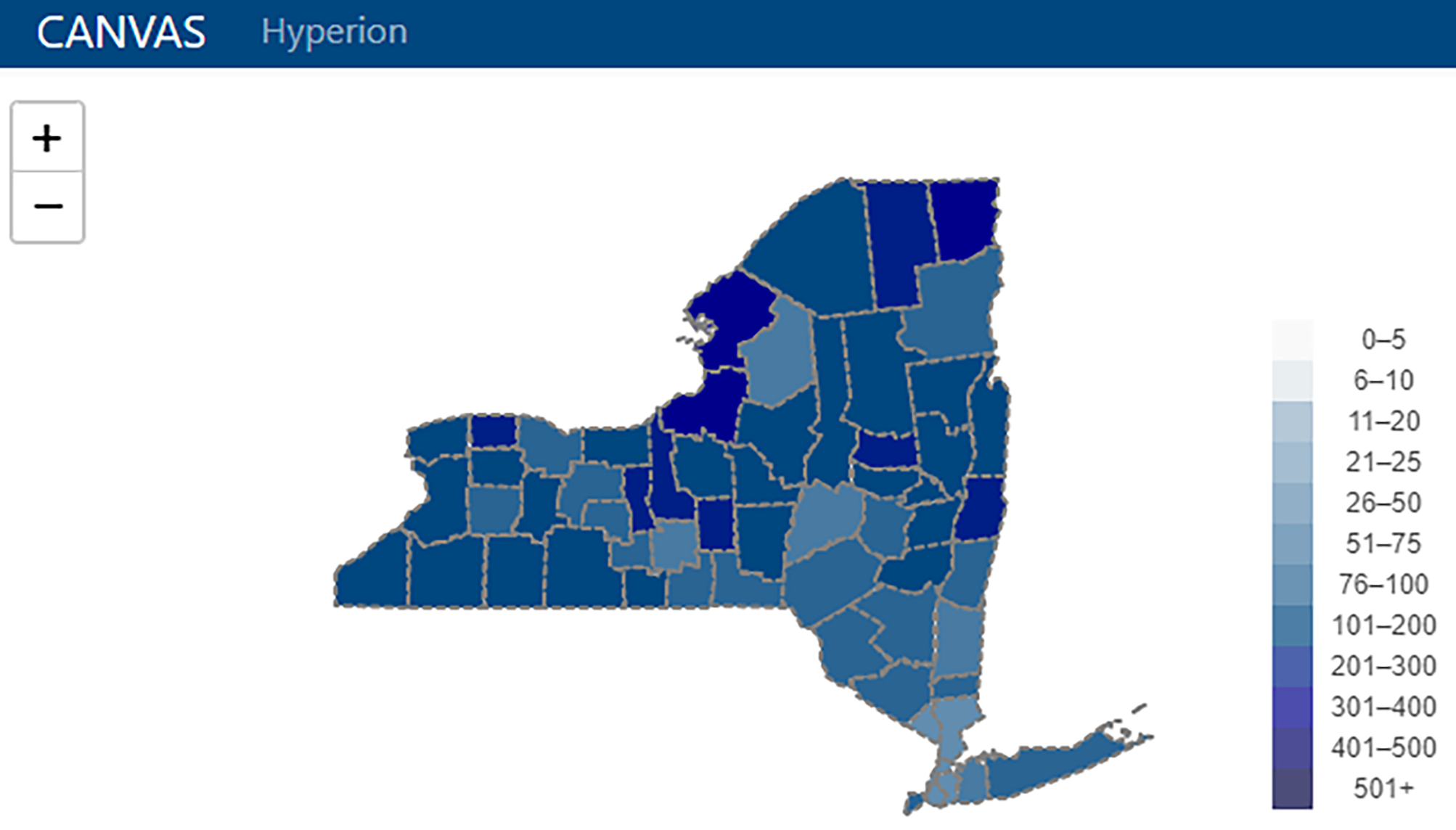
From disparate pools of data and information to inefficiencies in using that disparate data and information, the academic cancer center wishing to operate more effectively while improving patient outcomes and enabling more impactful research must turn to well-implemented data management systems and integrations. The University of Rochester Medical Center's James P. Wilmot Cancer Institute (WCI) was no exception to this greater need, turning to developing a custom data management platform called Hyperion to meet their needs. In this 2022 paper published in
PLOS Digital Health, key stakeholders of Hyperion at WCI present the background on cancer center data management, as well as their methods in developing and results in using Hyperion. They follow this with discussion of their results, concluding that "Hyperion has surmounted large challenges in working with healthcare data to merge, organize, validate, and package data for use in multiple applications" at WCI, while also "lowering the skill floor for interaction with and maintenance of the software, reducing costs, and encouraging user autonomy." They add that a similar such system, from their viewpoint, could provide similar benefits in other academic cancer centers.
Posted on February 13, 2023
By LabLynx
Journal articles
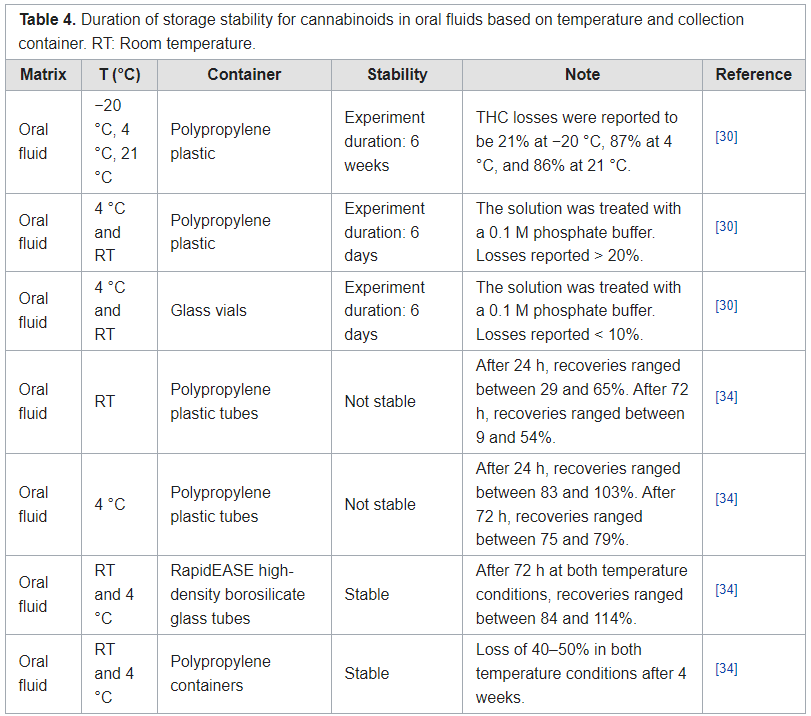
In this 2022 journal article published in
Metabolites, Djilali
et al. present the results of a literature review "of studies performed on the effects of different storage conditions on the stability of cannabis compounds present in various biological matrices." Noting a growing need for accurate testing for the presence of cannabis compounds in humans, the authors first review the state of analysis among conventional biological matrices (i.e., blood, plasma, urine, and oral fluids) before looking at alternative matrices (i.e., breath, bile fluid, hair, sweat, cerumen, and dried blood spots). From their literature analysis, the authors made several conclusions about the various matrices, their collection methods, and their testing. Depending on the matrix, the authors recommended specific sample containers and storage temperatures among the conventional matrices, while noting that more research is required concerning laboratory testing of alternative matrices.
Posted on February 7, 2023
By LabLynx
Journal articles
In this 2022 article published in the journal
BMC Medical Ethics, Scheibner
et al. present an assessment of Swiss hospital and medical research experts' views on using advanced technical solutions towards "sharing patient data in a privacy-preserving manner." In particular, the authors examine homomorphic encryption (HE) and distributed ledger technology (DLT) in the scope of European and Swiss regulatory frameworks. After an in-depth introduction on the topic, the authors present their methodology (an interview study) and their results, which touch upon five different information request scenarios. They then discuss the legal, ethical, and compatibility issues associated with using HE and DLT towards the problem of privacy-preserving patient data. They conclude that "a holistic approach needs to be taken to introducing HE and DLT as a mechanism for patient data management," one that tends "to recognize that social license and public trust from patients and physicians is as important as legal compliance."
Posted on January 31, 2023
By LabLynx
Journal articles
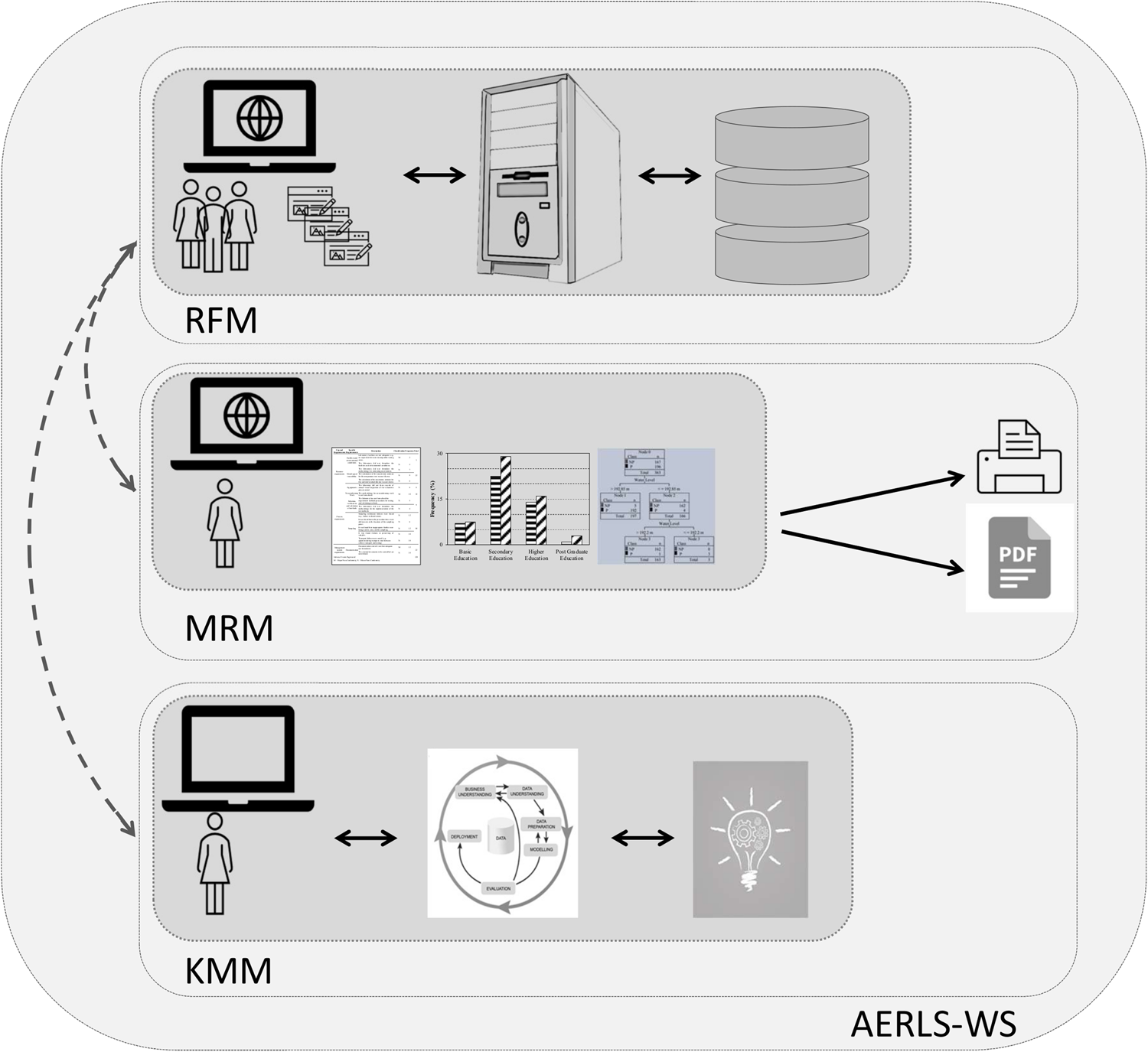
This 2022 journal article published in
AQUA - Water infrastructure, Ecosystems and Society sees Fernandes
et al. presenting a computerized "system for reporting and learning from adverse events" in water sampling and analysis laboratories. The work was prompted by "a high frequency of adverse events in connection with sampling" at the Water Laboratory of Santiago do Cacém Municipality in Portugal. After introducing the topic of water quality testing and work related to using artificial intelligence (AI) in that testing, the authors describe their method of using a Eindhoven Classification Model (ECM) within the scope of ISO/IEC 17025 requirements. They then discuss the results of their software-based approach and conclude that their modular adverse event reporting and learning system has many strengths, particularly in that it "allows knowledge extraction, i.e., the identification of the main failure causes, possible trends, areas requiring improvement plans, or changes in procedures."
Posted on January 23, 2023
By LabLynx
Journal articles
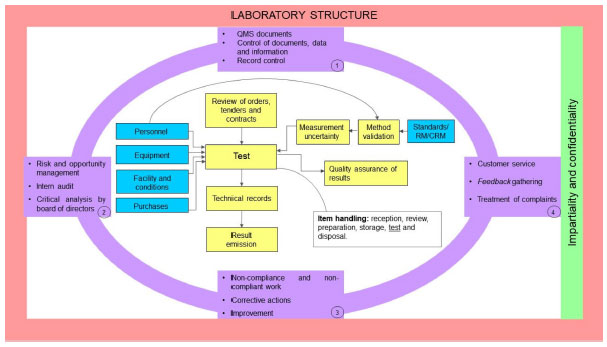
In this 2021 paper publisher in the journal
Química Nova, Miguel
et al. present the ISO/IEC 17025 standard in a historical context, while also providing some additional details about the concepts the standard proposes. As a quality management standard that also addresses competence, impartiality, and consistent operations in the laboratory, the standard emphasizes the importance of well-planned quality assurance towards reliable and traceable laboratory results. After an introduction to the standard, the authors take an in-depth look at ISO/IEC 17025's history and what drove changes to the standard over time. The article then examines what the standard asks of laboratories, as well as what value it adds to those labs. "By enacting ISO/IEC 17025," the authors conclude, "testing and calibration laboratories demonstrate they are responsible with their activities and their impacts, and put quality management and metrological traceability at the forefront of their operations."
Posted on January 16, 2023
By LabLynx
Journal articles
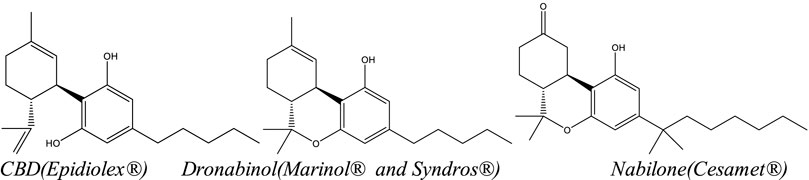
Hemp? Cannabis? Marijuana? Industrial hemp? Salehi
et al. note that terms like these and others are "a significant source of confusion for many," not just in the public but also within industry, marketing, and other groups. This not only causes issues with communicating research clearly but also employing that research to best effect. As such, the authors of this paper, published in the journal
Frontiers in Pharmacology, pose the important question, "What is essential for defining
Cannabis as a food, supplement, or drug?" Salehi
et al. answer this question by reviewing a wide swath of current study on the
Cannabis plant and its derivatives, highlighting critical definitions, composition, production practices, and pharmacological effects as part of defining the essentials of
Cannabis. The authors also go into its use in foods, drugs, and supplements, as well as regulatory status and testing methodologies. They conclude that "despite all of the recent advances, several cannabis topics remain to be addressed," and addressing those topics will require more consistent terminology, greater understanding of the human endocannabinoid system, and a more consistent "platform for developing research about cannabis and related industries" within a number of countries.
Posted on January 9, 2023
By LabLynx
Journal articles
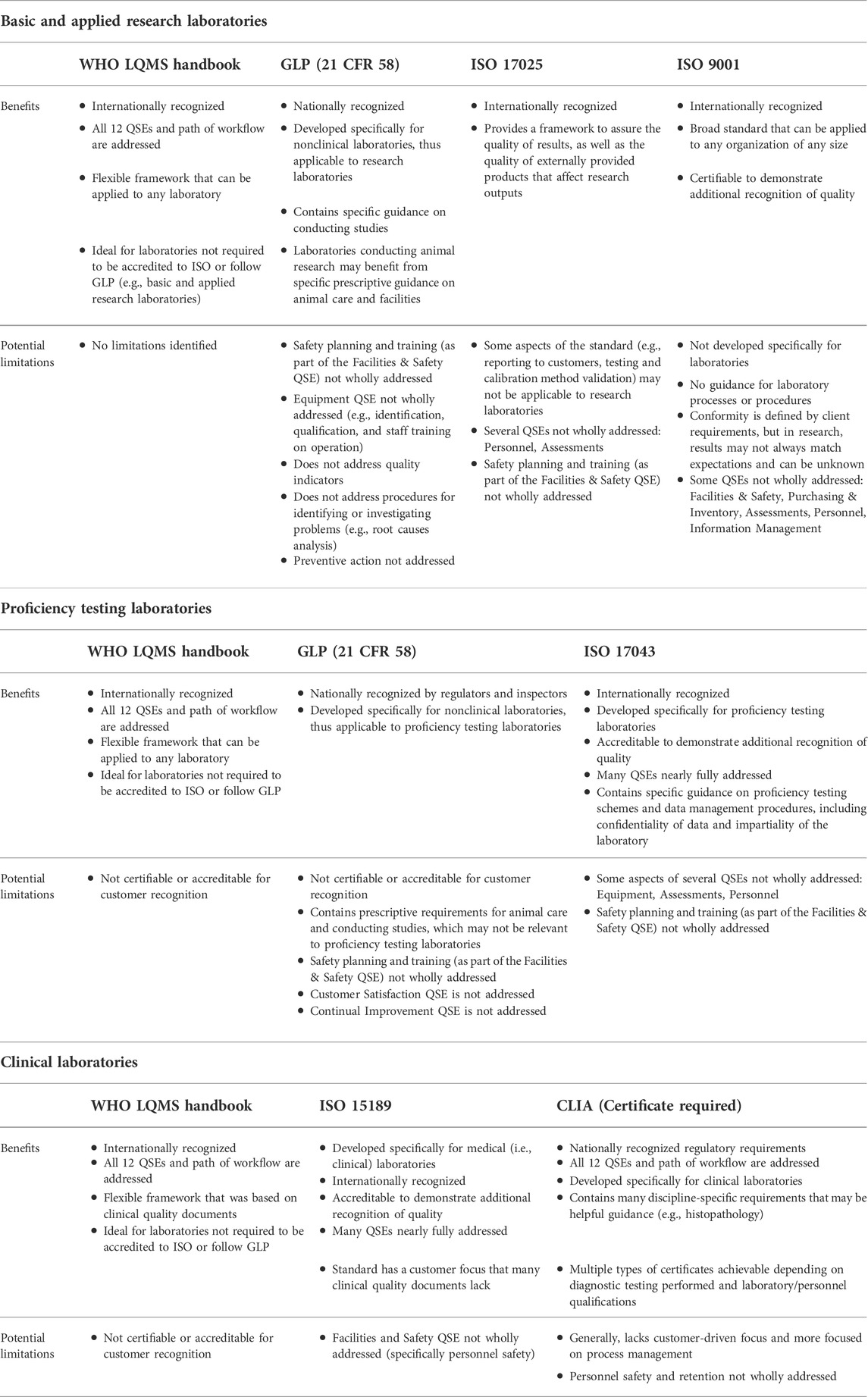
The quality management system (QMS) is increasingly a constant in laboratories of all types, lending support for better products and services coming out those labs. However, as technologies change and new challenges arise in research, medical, analytical, and R&D labs, the need for a more robust QMS becomes more obvious. In this 2022 article published in the journal
Frontiers in Bioengineering and Biotechnology, Pillai
et al. explore the varying frameworks of the QMS, which frameworks are best in which labs, and what implementation considerations should be made. After an informative introduction, the authors define the various laboratory types and match those types to the various QMS frameworks. They then provide a set of three critical recommendations for their implementation and use, concluding that while many frameworks are robust, the modern laboratory may need to incorporate more than one framework in order to fully achieve their quality goals. They add that "a holistic QMS framework complemented with guidance from multiple quality documents can benefit many laboratories that aim to address" the 12 quality system essentials (QSEs) of the Clinical and Laboratory Standards Institute (CLSI).
Posted on January 3, 2023
By LabLynx
Journal articles
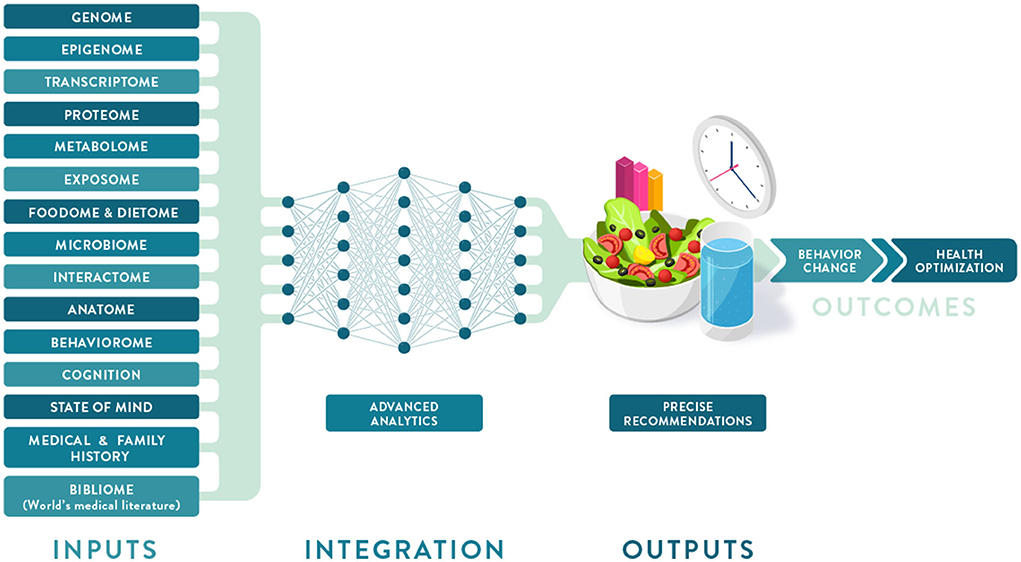
In this 2022 article published in the journal
Frontiers in Nutrition, Berciano
et al. discuss the topic of precision or personalized nutrition (PN) in the context of needing advancements in analytical and data management technologies in order to reach its full potential. After a brief introduction and discussion of the concept of PN, the authors examine the role of big data and data analysis in PN, particularly in regards to developing and optimizing personalized nutrition products and services. They then couch PN in terms of it having the potential to be "the future of healthcare," addressing topics such as phenotyping, metabolic methods, environmental factors, and the role of money and medical service insurers. After compiling a set of best practices and standards in PN and addressing the need for advocacy efforts towards PN, the authors conclude that while "PN should have an impact on both personal and public health ... [a]dvancing the science and the adoption of PN will require a significant investment in multidisciplinary collaborations that translate the fast-moving technological advances in omics, sensors, AI, and big data management and analytics into powerful and user-friendly tools."
Posted on December 19, 2022
By LabLynx
Journal articles
In this 2022 article published in the journal
Sustainability, da Silva
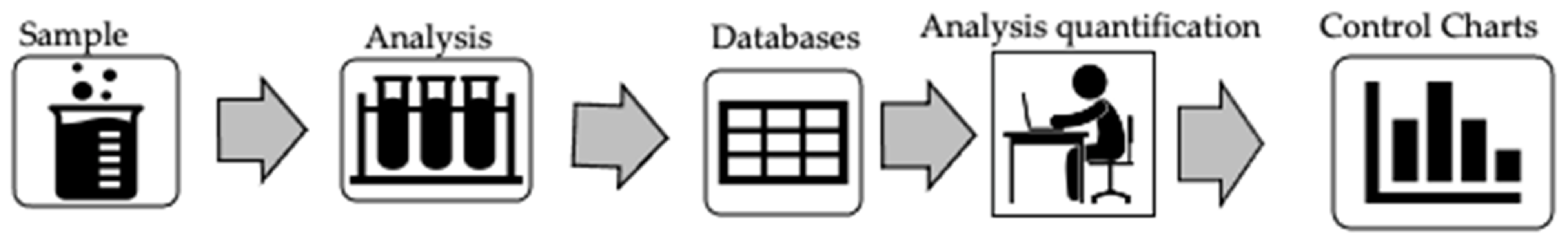
et al. of the Polytechnic Institute of Beja demonstrate the usefulness of control charting in the realm of maintaining water quality. Using a test case of wastewater from a treatment plant associated with a slaughterhouse, the authors describe their materials and methods, which progressed from wastewater sample to analysis and databasing of the results, followed by analysis quantification and control charting. In this article, they show 10 control charts associated with five test variables, demonstrating the usefulness of the control charts in identifying at least one significant set of out-of-specification results, likely associated with a treatment failure at the facility. They conclude that the results demonstrate "how rapid detection of variations in small scales on control charts allows the lab to better act upon identifying potential causes of variability," and they "show it is possible to state that all points are under statistical control, and thus by extension these types of control charts can have routine value in the laboratory."
Posted on December 13, 2022
By LabLynx
Journal articles

In this 2022 paper published in
Talanta Open, Hall
et al. propose an analytical method for 10 cannabis-related cannabinoids that is cost-efficient, rapid, and robust in its use. Noting "elevated cost and limited availability of certified analytical reference standards for some cannabinoids," the authors sought to develop a high-performance liquid chromatography-photodiode array (HPLC-PDA) method for 10 cannabinoids that uses "relative retention times (RRT) for peak identification and relative response factors (RRF) for their quantification." After describing their materials and methods, the authors present their results and discuss their implications towards various aspects such as accuracy, recovery, and quantification. They conclude that their method is robust, acceptably accurate, and has sufficient dynamic range. They add that with their method, "cost barriers for the analysis of panels of cannabinoids can be overcome, such that a diversity of cannabinoids can be analyzed as a part of routine quality control, with results that reflect the therapeutic efficacy for the consumer."
Posted on December 6, 2022
By LabLynx
Journal articles
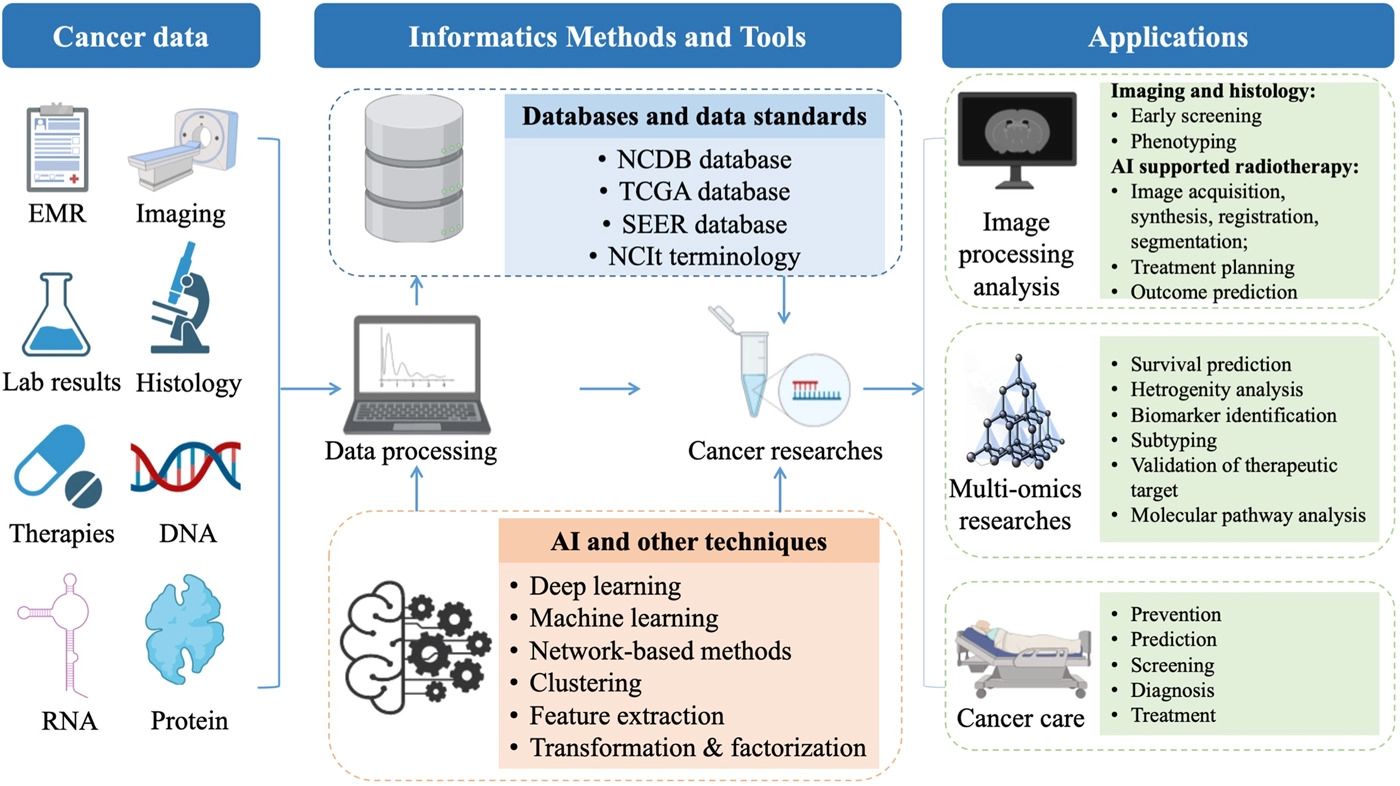
In the age of "big data," appropriately implemented informatics solutions have become more important than ever. This has become particularly important to the world of cancer research and the weighty analyses of "the genome, epigenome, transcriptome, proteome, metabolome, and microbiome" of individuals. In this 2022 journal article by Hong
et al., a review of the current state of cancer informatics is presented, addressing not only current informatics-supported applications but also challenges, opportunities, and future likelihoods. The authors conclude that not only "clinical oncology and research are reaping the benefits of informatics," but also that "with the further development of convenient and intelligent tools, informatics will enable earlier cancer detection, more precise cancer treatment, and better outcomes."
 In this 2023 article published in the journal Processes, Frede et al. of TU Dortmund University work with analytical consultancy d-fine GmbH to demonstrate experimental workflows of a microscale reaction calorimeter over d-fine's open-source internet of things (IoT) software platform. Their goal: provide an example of how data collection and capture methods can and should be standardized across an automated laboratory. After a brief introduction, the authors present their materials and methods, along with a case study of hydrolysis of acetic anhydride. After presenting the results of their case study, they conclude that "[t]he modular data platform complements the reaction calorimeter very well with additional functionalities," and by adding even more instruments, "[t]his approach will further harmonize data management within our laboratory while better applying FAIR Guiding Principles."
In this 2023 article published in the journal Processes, Frede et al. of TU Dortmund University work with analytical consultancy d-fine GmbH to demonstrate experimental workflows of a microscale reaction calorimeter over d-fine's open-source internet of things (IoT) software platform. Their goal: provide an example of how data collection and capture methods can and should be standardized across an automated laboratory. After a brief introduction, the authors present their materials and methods, along with a case study of hydrolysis of acetic anhydride. After presenting the results of their case study, they conclude that "[t]he modular data platform complements the reaction calorimeter very well with additional functionalities," and by adding even more instruments, "[t]his approach will further harmonize data management within our laboratory while better applying FAIR Guiding Principles."
 In this 2023 paper published in the journal Frontiers in Plant Science, Fernández et al. propose that Cannabis should be characterized "according to its chemical composition (i.e., its metabolome) and not only its botanical traits," while conducting a series of experiments to back their claim. After an introduction to cannabis and its chemical characterization, the authors describe the materials, methods, and results of their experimentation using metabolomics on high-THCA and high-CBDA C. sativa chemovars with medicinal potential. They conclude that many "differences between the two varieties beyond their cannabinoid composition" could be found through this experimentation, and that these differences could be "instrumental to breeders in the development of plant varieties agronomically adapted to specific environmental conditions." They also noted that fungal infections changed the metabolome of tested plants, which can better aid in the "classification of healthy and diseased plants."
In this 2023 paper published in the journal Frontiers in Plant Science, Fernández et al. propose that Cannabis should be characterized "according to its chemical composition (i.e., its metabolome) and not only its botanical traits," while conducting a series of experiments to back their claim. After an introduction to cannabis and its chemical characterization, the authors describe the materials, methods, and results of their experimentation using metabolomics on high-THCA and high-CBDA C. sativa chemovars with medicinal potential. They conclude that many "differences between the two varieties beyond their cannabinoid composition" could be found through this experimentation, and that these differences could be "instrumental to breeders in the development of plant varieties agronomically adapted to specific environmental conditions." They also noted that fungal infections changed the metabolome of tested plants, which can better aid in the "classification of healthy and diseased plants."
 In this 2022 paper published in Insights into Imaging, Beauchamp et al. focus in on the promise of "integrative diagnostics" as a means towards aggregating diagnostic data for more meaningful interpretation and contextualization, by extension translating to directly relatable clinical action. Looking specifically at radiology and pathology workflows, the authors describe what integrative diagnostics looks like, how the process should work, and what promise it holds for in vivo and in vitro radiology and pathology workflows. The authors turn to 24 different sources in highlighting the clinical potential of integrative diagnostics for a wide variety of diseases, highlighting the added clinical value and improved discovery possible with integrative diagnostics' application. After examining the current state of the art, the authors conclude by making several recommendations towards implementing integrative diagnostics, while also noting that "[c]reating financial models that demonstrate the economic value proposition of [integrative diagnostics] will be a necessary catalyst for change" for the overall health care system.
In this 2022 paper published in Insights into Imaging, Beauchamp et al. focus in on the promise of "integrative diagnostics" as a means towards aggregating diagnostic data for more meaningful interpretation and contextualization, by extension translating to directly relatable clinical action. Looking specifically at radiology and pathology workflows, the authors describe what integrative diagnostics looks like, how the process should work, and what promise it holds for in vivo and in vitro radiology and pathology workflows. The authors turn to 24 different sources in highlighting the clinical potential of integrative diagnostics for a wide variety of diseases, highlighting the added clinical value and improved discovery possible with integrative diagnostics' application. After examining the current state of the art, the authors conclude by making several recommendations towards implementing integrative diagnostics, while also noting that "[c]reating financial models that demonstrate the economic value proposition of [integrative diagnostics] will be a necessary catalyst for change" for the overall health care system.
 In January 2023, the National Institutes of Health's (NIH's) Policy for Data Management and Sharing came into effect, requiring guidance on elements "for the submission of a data management and sharing plan (DMSP)" for NIH-funded projects. While the NIH created supplementary guidance to developing a DMSP, some ambiguity arguably remained. In this 2022 article published in PLOS Computational Biology, Gonzales et al. provide additional insight into the process. After a brief introduction, the authors present 10 helpful tips towards getting the most of a DMSP in complying with the NIH and its requirements. They conclude by examining the various stakeholders involved with the DMSP process and finding that "[g]ood DMSP practices can support better engagement with and accountability to the public who benefit from research."
In January 2023, the National Institutes of Health's (NIH's) Policy for Data Management and Sharing came into effect, requiring guidance on elements "for the submission of a data management and sharing plan (DMSP)" for NIH-funded projects. While the NIH created supplementary guidance to developing a DMSP, some ambiguity arguably remained. In this 2022 article published in PLOS Computational Biology, Gonzales et al. provide additional insight into the process. After a brief introduction, the authors present 10 helpful tips towards getting the most of a DMSP in complying with the NIH and its requirements. They conclude by examining the various stakeholders involved with the DMSP process and finding that "[g]ood DMSP practices can support better engagement with and accountability to the public who benefit from research."
 In this 2022 journal article published in Electronics, University Canada West's Hamed Taherdoost presents a narrative literature review of a variety of cybersecurity standards and frameworks as a means to "help organizations select the cybersecurity standard or framework that best fits their cybersecurity requirements." After providing a brief introduction, Taherdoost speaks generally about cybersecurity standards and frameworks, and what they help organizations of all types achieve with their own cybersecurity. The author then goes into greater detail of those standards and frameworks, as well as their supporting documentation. Taherdoost then presents the methodology for his literature review, followed by a discussion of the results of that review. The author concludes that the presented standards and frameworks help organizations "ensure the security of data against cyber threats," though one standard or framework "may not fulfill all the demands of an organization, and it may be necessary to employ a combination of standards in order to ensure security against cyber threats and data loss."
In this 2022 journal article published in Electronics, University Canada West's Hamed Taherdoost presents a narrative literature review of a variety of cybersecurity standards and frameworks as a means to "help organizations select the cybersecurity standard or framework that best fits their cybersecurity requirements." After providing a brief introduction, Taherdoost speaks generally about cybersecurity standards and frameworks, and what they help organizations of all types achieve with their own cybersecurity. The author then goes into greater detail of those standards and frameworks, as well as their supporting documentation. Taherdoost then presents the methodology for his literature review, followed by a discussion of the results of that review. The author concludes that the presented standards and frameworks help organizations "ensure the security of data against cyber threats," though one standard or framework "may not fulfill all the demands of an organization, and it may be necessary to employ a combination of standards in order to ensure security against cyber threats and data loss."
 In this 2022 journal article published in Journal of Integrative Bioinformatics, Panse et al. discuss the state of research-based core facilities that share research resources across institutions or organizations, in particular in regards to data and information management. The authors note that the "interdisciplinary nature of such scientific projects requires a powerful and flexible platform for data annotation, communication, and exploration," and the authors—associated with the Functional Genomics Center Zurich (FGCZ)—present B-Fabric, their solution to data and information management in such settings. After providing a brief background, the authors discuss the development and application of B-Fabric at FGCZ, along with lessons learned. They conclude that with systems like B-Fabric, "ad-hoc visualizations and analytics can be realized on top of an integrative platform that captures and processes any kind of data and scientific processes." The platform is also able "to decouple metadata management from data processing and visualization components" while also optimizing specific software software environments for different research areas.
In this 2022 journal article published in Journal of Integrative Bioinformatics, Panse et al. discuss the state of research-based core facilities that share research resources across institutions or organizations, in particular in regards to data and information management. The authors note that the "interdisciplinary nature of such scientific projects requires a powerful and flexible platform for data annotation, communication, and exploration," and the authors—associated with the Functional Genomics Center Zurich (FGCZ)—present B-Fabric, their solution to data and information management in such settings. After providing a brief background, the authors discuss the development and application of B-Fabric at FGCZ, along with lessons learned. They conclude that with systems like B-Fabric, "ad-hoc visualizations and analytics can be realized on top of an integrative platform that captures and processes any kind of data and scientific processes." The platform is also able "to decouple metadata management from data processing and visualization components" while also optimizing specific software software environments for different research areas.
 Genotyping of Cannabis sativa is a useful component of not only gender identification of plants for cultivators but also other types of laboratory testing and research of Cannabis. Performing such tests in a reliable, cost-effective, and high-performance way has significant value for Cannabis testing laboratories. In this 2022 journal article, Torres et al. compare a number of different genotyping techniques, ultimately demonstrating some of that value. After providing background information and their methodology, the authors discuss the results of gender testing using a wide array of genotyping techniques. They conclude that "the multiple methodologies presented here allow for accurate, quick, and cost-effective screening that will enable future development of germplasm and the industry." They add that their "real-time assays can be performed by cannabis testing laboratories performing diagnostic testing, as well as in the field with minimal molecular biology equipment and expertise."
Genotyping of Cannabis sativa is a useful component of not only gender identification of plants for cultivators but also other types of laboratory testing and research of Cannabis. Performing such tests in a reliable, cost-effective, and high-performance way has significant value for Cannabis testing laboratories. In this 2022 journal article, Torres et al. compare a number of different genotyping techniques, ultimately demonstrating some of that value. After providing background information and their methodology, the authors discuss the results of gender testing using a wide array of genotyping techniques. They conclude that "the multiple methodologies presented here allow for accurate, quick, and cost-effective screening that will enable future development of germplasm and the industry." They add that their "real-time assays can be performed by cannabis testing laboratories performing diagnostic testing, as well as in the field with minimal molecular biology equipment and expertise."
 In this 2022 journal article published in Applied Sciences, Mustapää et al. turn to the calibration process of the pharmaceutical industry—as well as their regulatory- and standards-based requirements—as inspiration for building a proof-of-concept software platform that better ensures "the preservation of traceability and data integrity" of calibration data. After a lengthy and thorough background to the topic of laboratory calibrations and calibration data management, the authors present their proof of concept software platform, designed "to harmonize the data exchange between the parties of a calibration chain to enable automation of the data management processes in the receiving end." They then discuss the results of their efforts, challenges in advancing such metrology infrastructure, and topics that require further research. They conclude that the real use cases tested on their proof of concept "proved that the multitenant platform is an efficient means of organizing the exchange of [digital calibration certificates]," and that "the system would suit the requirements of other industries as well, providing cost-efficiency through economies of scale."
In this 2022 journal article published in Applied Sciences, Mustapää et al. turn to the calibration process of the pharmaceutical industry—as well as their regulatory- and standards-based requirements—as inspiration for building a proof-of-concept software platform that better ensures "the preservation of traceability and data integrity" of calibration data. After a lengthy and thorough background to the topic of laboratory calibrations and calibration data management, the authors present their proof of concept software platform, designed "to harmonize the data exchange between the parties of a calibration chain to enable automation of the data management processes in the receiving end." They then discuss the results of their efforts, challenges in advancing such metrology infrastructure, and topics that require further research. They conclude that the real use cases tested on their proof of concept "proved that the multitenant platform is an efficient means of organizing the exchange of [digital calibration certificates]," and that "the system would suit the requirements of other industries as well, providing cost-efficiency through economies of scale."
 In this 2022 paper published in the Egyptian Journal of Forensic Sciences, Prahladh and van Wyk provide a first-of-its-kind comprehensive scoping review of "articles involving unnatural deaths, focused on data practice or data management systems, relating to forensic medicine, all study designs, and published in English," with the goal of being able to draw conclusions about the current academic view of forensic data management. After examining more than 23,000 articles, the authors were only able to find 16 that met their criteria, according to their stated methodology. After a thorough review of those 16 articles, the duo concludes that "the literature shows that electronic data reporting systems are relevant and were developed from the recognition of coronial data as not only a part of the death investigation but as a contributor to preventable death research and public health initiatives." However, there are few lower- and middle-income countries (LMICs) represented in the study, and the authors believe that given how "forensic medicine departments can utilize simple and available tools that can advance standardization of data collection, storage, and reporting," it would behoove more LMICs to consider pilot systems such as "the Nigerian pilot program [that] still managed to contribute to statistically relevant data for impactful research" while using low-cost and readily available data management tools.
In this 2022 paper published in the Egyptian Journal of Forensic Sciences, Prahladh and van Wyk provide a first-of-its-kind comprehensive scoping review of "articles involving unnatural deaths, focused on data practice or data management systems, relating to forensic medicine, all study designs, and published in English," with the goal of being able to draw conclusions about the current academic view of forensic data management. After examining more than 23,000 articles, the authors were only able to find 16 that met their criteria, according to their stated methodology. After a thorough review of those 16 articles, the duo concludes that "the literature shows that electronic data reporting systems are relevant and were developed from the recognition of coronial data as not only a part of the death investigation but as a contributor to preventable death research and public health initiatives." However, there are few lower- and middle-income countries (LMICs) represented in the study, and the authors believe that given how "forensic medicine departments can utilize simple and available tools that can advance standardization of data collection, storage, and reporting," it would behoove more LMICs to consider pilot systems such as "the Nigerian pilot program [that] still managed to contribute to statistically relevant data for impactful research" while using low-cost and readily available data management tools.
 From disparate pools of data and information to inefficiencies in using that disparate data and information, the academic cancer center wishing to operate more effectively while improving patient outcomes and enabling more impactful research must turn to well-implemented data management systems and integrations. The University of Rochester Medical Center's James P. Wilmot Cancer Institute (WCI) was no exception to this greater need, turning to developing a custom data management platform called Hyperion to meet their needs. In this 2022 paper published in PLOS Digital Health, key stakeholders of Hyperion at WCI present the background on cancer center data management, as well as their methods in developing and results in using Hyperion. They follow this with discussion of their results, concluding that "Hyperion has surmounted large challenges in working with healthcare data to merge, organize, validate, and package data for use in multiple applications" at WCI, while also "lowering the skill floor for interaction with and maintenance of the software, reducing costs, and encouraging user autonomy." They add that a similar such system, from their viewpoint, could provide similar benefits in other academic cancer centers.
From disparate pools of data and information to inefficiencies in using that disparate data and information, the academic cancer center wishing to operate more effectively while improving patient outcomes and enabling more impactful research must turn to well-implemented data management systems and integrations. The University of Rochester Medical Center's James P. Wilmot Cancer Institute (WCI) was no exception to this greater need, turning to developing a custom data management platform called Hyperion to meet their needs. In this 2022 paper published in PLOS Digital Health, key stakeholders of Hyperion at WCI present the background on cancer center data management, as well as their methods in developing and results in using Hyperion. They follow this with discussion of their results, concluding that "Hyperion has surmounted large challenges in working with healthcare data to merge, organize, validate, and package data for use in multiple applications" at WCI, while also "lowering the skill floor for interaction with and maintenance of the software, reducing costs, and encouraging user autonomy." They add that a similar such system, from their viewpoint, could provide similar benefits in other academic cancer centers.
 In this 2022 journal article published in Metabolites, Djilali et al. present the results of a literature review "of studies performed on the effects of different storage conditions on the stability of cannabis compounds present in various biological matrices." Noting a growing need for accurate testing for the presence of cannabis compounds in humans, the authors first review the state of analysis among conventional biological matrices (i.e., blood, plasma, urine, and oral fluids) before looking at alternative matrices (i.e., breath, bile fluid, hair, sweat, cerumen, and dried blood spots). From their literature analysis, the authors made several conclusions about the various matrices, their collection methods, and their testing. Depending on the matrix, the authors recommended specific sample containers and storage temperatures among the conventional matrices, while noting that more research is required concerning laboratory testing of alternative matrices.
In this 2022 journal article published in Metabolites, Djilali et al. present the results of a literature review "of studies performed on the effects of different storage conditions on the stability of cannabis compounds present in various biological matrices." Noting a growing need for accurate testing for the presence of cannabis compounds in humans, the authors first review the state of analysis among conventional biological matrices (i.e., blood, plasma, urine, and oral fluids) before looking at alternative matrices (i.e., breath, bile fluid, hair, sweat, cerumen, and dried blood spots). From their literature analysis, the authors made several conclusions about the various matrices, their collection methods, and their testing. Depending on the matrix, the authors recommended specific sample containers and storage temperatures among the conventional matrices, while noting that more research is required concerning laboratory testing of alternative matrices.
 In this 2022 article published in the journal BMC Medical Ethics, Scheibner et al. present an assessment of Swiss hospital and medical research experts' views on using advanced technical solutions towards "sharing patient data in a privacy-preserving manner." In particular, the authors examine homomorphic encryption (HE) and distributed ledger technology (DLT) in the scope of European and Swiss regulatory frameworks. After an in-depth introduction on the topic, the authors present their methodology (an interview study) and their results, which touch upon five different information request scenarios. They then discuss the legal, ethical, and compatibility issues associated with using HE and DLT towards the problem of privacy-preserving patient data. They conclude that "a holistic approach needs to be taken to introducing HE and DLT as a mechanism for patient data management," one that tends "to recognize that social license and public trust from patients and physicians is as important as legal compliance."
In this 2022 article published in the journal BMC Medical Ethics, Scheibner et al. present an assessment of Swiss hospital and medical research experts' views on using advanced technical solutions towards "sharing patient data in a privacy-preserving manner." In particular, the authors examine homomorphic encryption (HE) and distributed ledger technology (DLT) in the scope of European and Swiss regulatory frameworks. After an in-depth introduction on the topic, the authors present their methodology (an interview study) and their results, which touch upon five different information request scenarios. They then discuss the legal, ethical, and compatibility issues associated with using HE and DLT towards the problem of privacy-preserving patient data. They conclude that "a holistic approach needs to be taken to introducing HE and DLT as a mechanism for patient data management," one that tends "to recognize that social license and public trust from patients and physicians is as important as legal compliance."
 This 2022 journal article published in AQUA - Water infrastructure, Ecosystems and Society sees Fernandes et al. presenting a computerized "system for reporting and learning from adverse events" in water sampling and analysis laboratories. The work was prompted by "a high frequency of adverse events in connection with sampling" at the Water Laboratory of Santiago do Cacém Municipality in Portugal. After introducing the topic of water quality testing and work related to using artificial intelligence (AI) in that testing, the authors describe their method of using a Eindhoven Classification Model (ECM) within the scope of ISO/IEC 17025 requirements. They then discuss the results of their software-based approach and conclude that their modular adverse event reporting and learning system has many strengths, particularly in that it "allows knowledge extraction, i.e., the identification of the main failure causes, possible trends, areas requiring improvement plans, or changes in procedures."
This 2022 journal article published in AQUA - Water infrastructure, Ecosystems and Society sees Fernandes et al. presenting a computerized "system for reporting and learning from adverse events" in water sampling and analysis laboratories. The work was prompted by "a high frequency of adverse events in connection with sampling" at the Water Laboratory of Santiago do Cacém Municipality in Portugal. After introducing the topic of water quality testing and work related to using artificial intelligence (AI) in that testing, the authors describe their method of using a Eindhoven Classification Model (ECM) within the scope of ISO/IEC 17025 requirements. They then discuss the results of their software-based approach and conclude that their modular adverse event reporting and learning system has many strengths, particularly in that it "allows knowledge extraction, i.e., the identification of the main failure causes, possible trends, areas requiring improvement plans, or changes in procedures."
 In this 2021 paper publisher in the journal Química Nova, Miguel et al. present the ISO/IEC 17025 standard in a historical context, while also providing some additional details about the concepts the standard proposes. As a quality management standard that also addresses competence, impartiality, and consistent operations in the laboratory, the standard emphasizes the importance of well-planned quality assurance towards reliable and traceable laboratory results. After an introduction to the standard, the authors take an in-depth look at ISO/IEC 17025's history and what drove changes to the standard over time. The article then examines what the standard asks of laboratories, as well as what value it adds to those labs. "By enacting ISO/IEC 17025," the authors conclude, "testing and calibration laboratories demonstrate they are responsible with their activities and their impacts, and put quality management and metrological traceability at the forefront of their operations."
In this 2021 paper publisher in the journal Química Nova, Miguel et al. present the ISO/IEC 17025 standard in a historical context, while also providing some additional details about the concepts the standard proposes. As a quality management standard that also addresses competence, impartiality, and consistent operations in the laboratory, the standard emphasizes the importance of well-planned quality assurance towards reliable and traceable laboratory results. After an introduction to the standard, the authors take an in-depth look at ISO/IEC 17025's history and what drove changes to the standard over time. The article then examines what the standard asks of laboratories, as well as what value it adds to those labs. "By enacting ISO/IEC 17025," the authors conclude, "testing and calibration laboratories demonstrate they are responsible with their activities and their impacts, and put quality management and metrological traceability at the forefront of their operations."
 Hemp? Cannabis? Marijuana? Industrial hemp? Salehi et al. note that terms like these and others are "a significant source of confusion for many," not just in the public but also within industry, marketing, and other groups. This not only causes issues with communicating research clearly but also employing that research to best effect. As such, the authors of this paper, published in the journal Frontiers in Pharmacology, pose the important question, "What is essential for defining Cannabis as a food, supplement, or drug?" Salehi et al. answer this question by reviewing a wide swath of current study on the Cannabis plant and its derivatives, highlighting critical definitions, composition, production practices, and pharmacological effects as part of defining the essentials of Cannabis. The authors also go into its use in foods, drugs, and supplements, as well as regulatory status and testing methodologies. They conclude that "despite all of the recent advances, several cannabis topics remain to be addressed," and addressing those topics will require more consistent terminology, greater understanding of the human endocannabinoid system, and a more consistent "platform for developing research about cannabis and related industries" within a number of countries.
Hemp? Cannabis? Marijuana? Industrial hemp? Salehi et al. note that terms like these and others are "a significant source of confusion for many," not just in the public but also within industry, marketing, and other groups. This not only causes issues with communicating research clearly but also employing that research to best effect. As such, the authors of this paper, published in the journal Frontiers in Pharmacology, pose the important question, "What is essential for defining Cannabis as a food, supplement, or drug?" Salehi et al. answer this question by reviewing a wide swath of current study on the Cannabis plant and its derivatives, highlighting critical definitions, composition, production practices, and pharmacological effects as part of defining the essentials of Cannabis. The authors also go into its use in foods, drugs, and supplements, as well as regulatory status and testing methodologies. They conclude that "despite all of the recent advances, several cannabis topics remain to be addressed," and addressing those topics will require more consistent terminology, greater understanding of the human endocannabinoid system, and a more consistent "platform for developing research about cannabis and related industries" within a number of countries.
 The quality management system (QMS) is increasingly a constant in laboratories of all types, lending support for better products and services coming out those labs. However, as technologies change and new challenges arise in research, medical, analytical, and R&D labs, the need for a more robust QMS becomes more obvious. In this 2022 article published in the journal Frontiers in Bioengineering and Biotechnology, Pillai et al. explore the varying frameworks of the QMS, which frameworks are best in which labs, and what implementation considerations should be made. After an informative introduction, the authors define the various laboratory types and match those types to the various QMS frameworks. They then provide a set of three critical recommendations for their implementation and use, concluding that while many frameworks are robust, the modern laboratory may need to incorporate more than one framework in order to fully achieve their quality goals. They add that "a holistic QMS framework complemented with guidance from multiple quality documents can benefit many laboratories that aim to address" the 12 quality system essentials (QSEs) of the Clinical and Laboratory Standards Institute (CLSI).
The quality management system (QMS) is increasingly a constant in laboratories of all types, lending support for better products and services coming out those labs. However, as technologies change and new challenges arise in research, medical, analytical, and R&D labs, the need for a more robust QMS becomes more obvious. In this 2022 article published in the journal Frontiers in Bioengineering and Biotechnology, Pillai et al. explore the varying frameworks of the QMS, which frameworks are best in which labs, and what implementation considerations should be made. After an informative introduction, the authors define the various laboratory types and match those types to the various QMS frameworks. They then provide a set of three critical recommendations for their implementation and use, concluding that while many frameworks are robust, the modern laboratory may need to incorporate more than one framework in order to fully achieve their quality goals. They add that "a holistic QMS framework complemented with guidance from multiple quality documents can benefit many laboratories that aim to address" the 12 quality system essentials (QSEs) of the Clinical and Laboratory Standards Institute (CLSI).
 In this 2022 article published in the journal Frontiers in Nutrition, Berciano et al. discuss the topic of precision or personalized nutrition (PN) in the context of needing advancements in analytical and data management technologies in order to reach its full potential. After a brief introduction and discussion of the concept of PN, the authors examine the role of big data and data analysis in PN, particularly in regards to developing and optimizing personalized nutrition products and services. They then couch PN in terms of it having the potential to be "the future of healthcare," addressing topics such as phenotyping, metabolic methods, environmental factors, and the role of money and medical service insurers. After compiling a set of best practices and standards in PN and addressing the need for advocacy efforts towards PN, the authors conclude that while "PN should have an impact on both personal and public health ... [a]dvancing the science and the adoption of PN will require a significant investment in multidisciplinary collaborations that translate the fast-moving technological advances in omics, sensors, AI, and big data management and analytics into powerful and user-friendly tools."
In this 2022 article published in the journal Frontiers in Nutrition, Berciano et al. discuss the topic of precision or personalized nutrition (PN) in the context of needing advancements in analytical and data management technologies in order to reach its full potential. After a brief introduction and discussion of the concept of PN, the authors examine the role of big data and data analysis in PN, particularly in regards to developing and optimizing personalized nutrition products and services. They then couch PN in terms of it having the potential to be "the future of healthcare," addressing topics such as phenotyping, metabolic methods, environmental factors, and the role of money and medical service insurers. After compiling a set of best practices and standards in PN and addressing the need for advocacy efforts towards PN, the authors conclude that while "PN should have an impact on both personal and public health ... [a]dvancing the science and the adoption of PN will require a significant investment in multidisciplinary collaborations that translate the fast-moving technological advances in omics, sensors, AI, and big data management and analytics into powerful and user-friendly tools."
 In this 2022 paper published in Talanta Open, Hall et al. propose an analytical method for 10 cannabis-related cannabinoids that is cost-efficient, rapid, and robust in its use. Noting "elevated cost and limited availability of certified analytical reference standards for some cannabinoids," the authors sought to develop a high-performance liquid chromatography-photodiode array (HPLC-PDA) method for 10 cannabinoids that uses "relative retention times (RRT) for peak identification and relative response factors (RRF) for their quantification." After describing their materials and methods, the authors present their results and discuss their implications towards various aspects such as accuracy, recovery, and quantification. They conclude that their method is robust, acceptably accurate, and has sufficient dynamic range. They add that with their method, "cost barriers for the analysis of panels of cannabinoids can be overcome, such that a diversity of cannabinoids can be analyzed as a part of routine quality control, with results that reflect the therapeutic efficacy for the consumer."
In this 2022 paper published in Talanta Open, Hall et al. propose an analytical method for 10 cannabis-related cannabinoids that is cost-efficient, rapid, and robust in its use. Noting "elevated cost and limited availability of certified analytical reference standards for some cannabinoids," the authors sought to develop a high-performance liquid chromatography-photodiode array (HPLC-PDA) method for 10 cannabinoids that uses "relative retention times (RRT) for peak identification and relative response factors (RRF) for their quantification." After describing their materials and methods, the authors present their results and discuss their implications towards various aspects such as accuracy, recovery, and quantification. They conclude that their method is robust, acceptably accurate, and has sufficient dynamic range. They add that with their method, "cost barriers for the analysis of panels of cannabinoids can be overcome, such that a diversity of cannabinoids can be analyzed as a part of routine quality control, with results that reflect the therapeutic efficacy for the consumer."
 In the age of "big data," appropriately implemented informatics solutions have become more important than ever. This has become particularly important to the world of cancer research and the weighty analyses of "the genome, epigenome, transcriptome, proteome, metabolome, and microbiome" of individuals. In this 2022 journal article by Hong et al., a review of the current state of cancer informatics is presented, addressing not only current informatics-supported applications but also challenges, opportunities, and future likelihoods. The authors conclude that not only "clinical oncology and research are reaping the benefits of informatics," but also that "with the further development of convenient and intelligent tools, informatics will enable earlier cancer detection, more precise cancer treatment, and better outcomes."
In the age of "big data," appropriately implemented informatics solutions have become more important than ever. This has become particularly important to the world of cancer research and the weighty analyses of "the genome, epigenome, transcriptome, proteome, metabolome, and microbiome" of individuals. In this 2022 journal article by Hong et al., a review of the current state of cancer informatics is presented, addressing not only current informatics-supported applications but also challenges, opportunities, and future likelihoods. The authors conclude that not only "clinical oncology and research are reaping the benefits of informatics," but also that "with the further development of convenient and intelligent tools, informatics will enable earlier cancer detection, more precise cancer treatment, and better outcomes."







