Posted on August 11, 2020
By LabLynx
Journal articles
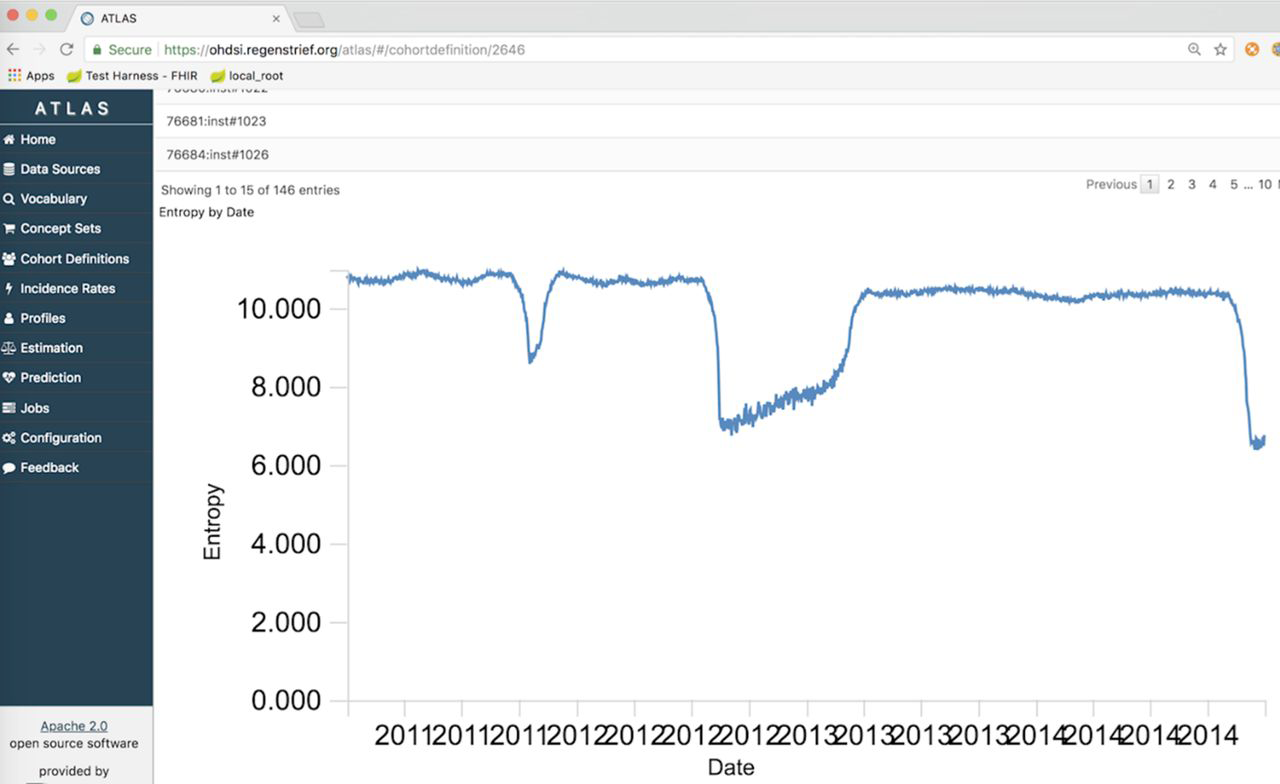
In this 2020 short report from Dixon
et al., lessons learned from attempts to expand data quality measures in the open-source tool Observational Health Data Science and Informatics (OHDSI) are presented. Noting a lack of data quality assessment and improvement mechanisms in health information systems in general, the researchers sought to improve OHDSI for public health surveillance use cases. After explaining the practical uses of OHDSI, the authors state their case for why measuring completeness, timeliness, and information entropy within OHDSI would be useful. Though not getting approval to add timeliness to the system, they conclude that high value remains "in adapting existing infrastructure and tools to support expanded use cases rather than to just create independent tools for use by a niche group."
Posted on August 4, 2020
By LabLynx
Journal articles
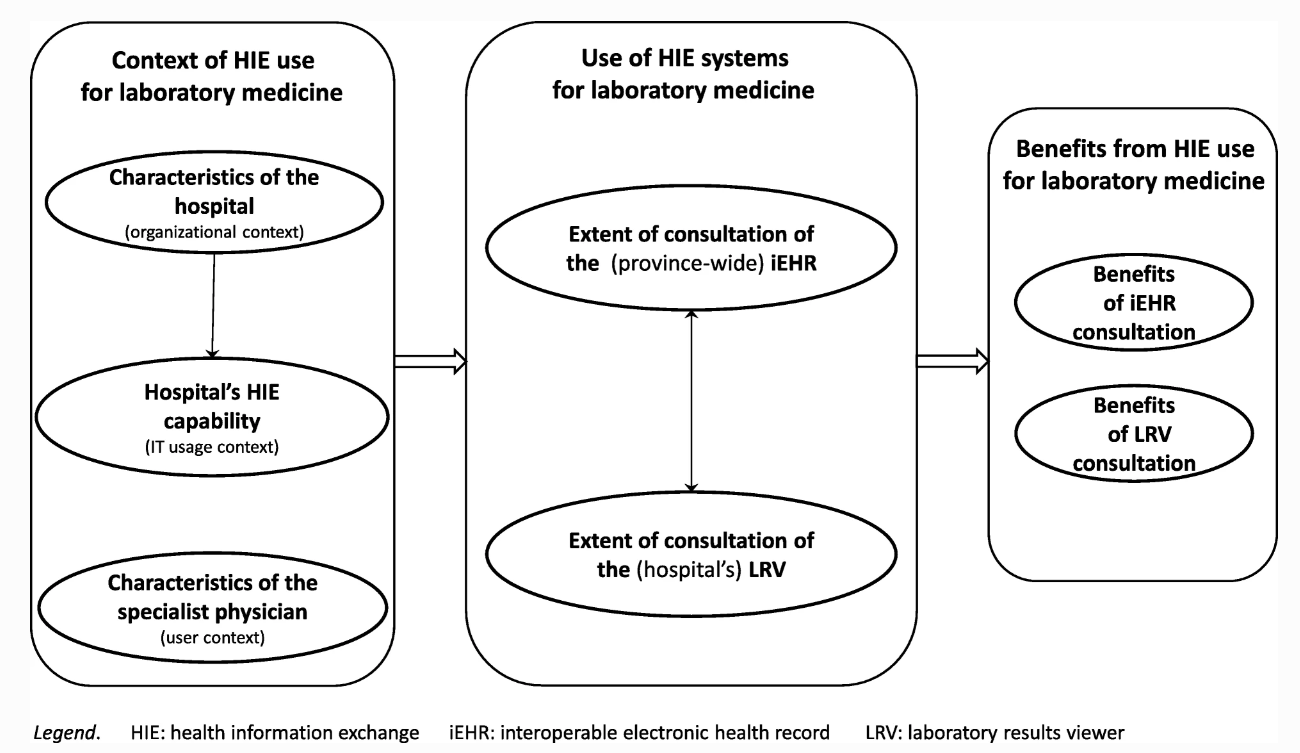
This February 2020 paper published in
BMC Medical Informatics and Decision Making examines the state of health information exchange in Québec and other parts of Canada and how its application to laboratory medicine might be improved. In particular, laboratory information exchange (LIE) systems that "improve the reliability of the laboratory testing process" and integrate "with other clinical information systems (CISs) physicians use in hospitals" are examined in this work. Surveying hospital-based specialist physicians, Raymond
et al. paint a picture of how varying clinical information management solutions are used, what functionality is being used and not used, and how physicians view the potential benefits of the clinical systems they use. They conclude that there is very much a "complementary nature" between systems and that "system designers should take a step back to imagine a way to design systems as part of an interconnected network of features."
Posted on July 28, 2020
By LabLynx
Journal articles
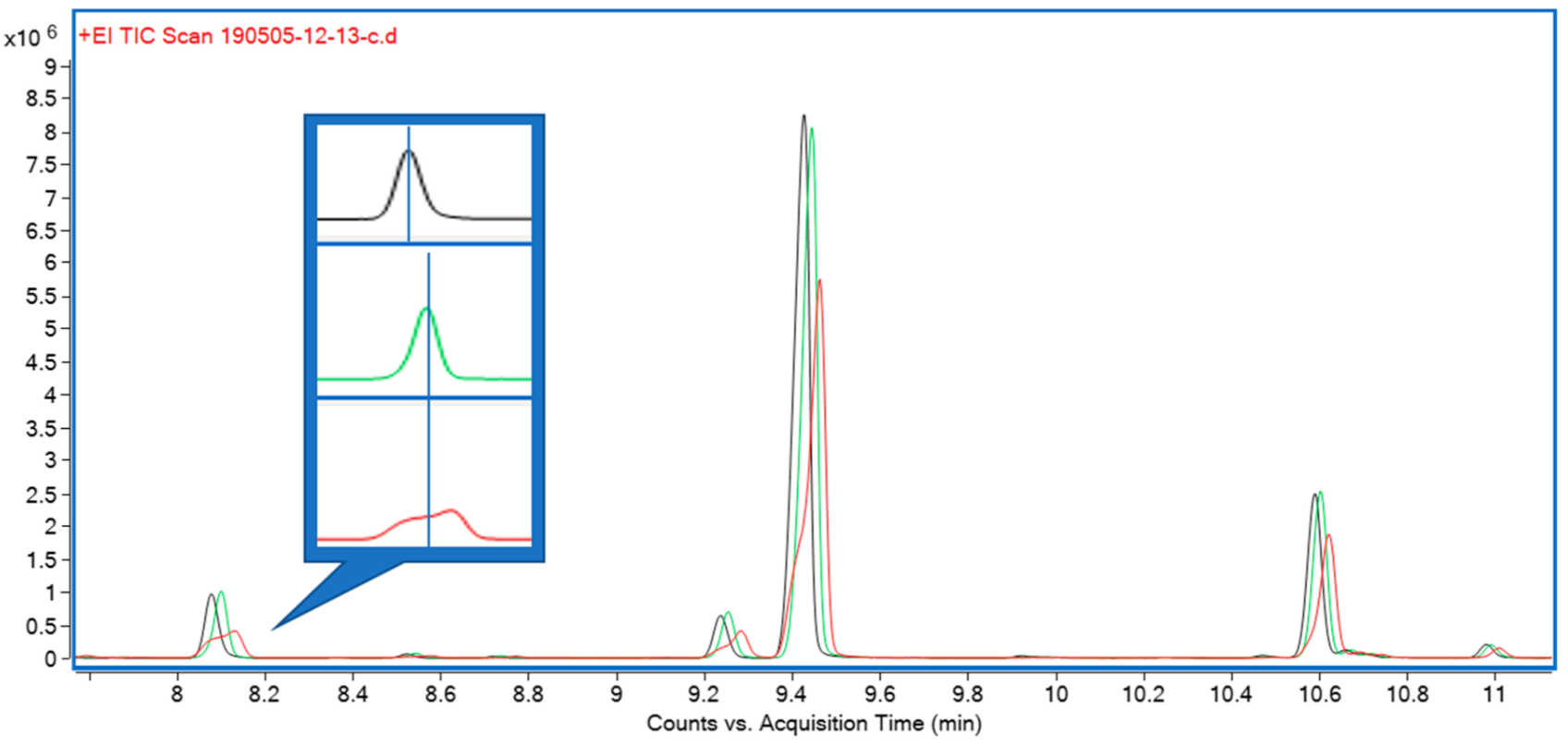
In this 2020 paper published in the journal
Matabolites, Krill
et al. of Australia's AgriBio present their findings in an attempt to reduce runtimes and extraction complexity associated with quantifying terpenes in cannabis biomass. Noting the evolving needs of high-throughput cannabis breeding programs, the researchers present a method "based on a simple hexane extract from 40 mg of biomass, with 50 μg/mL dodecane as internal standard, and a gradient of less than 30 minutes." After presenting current background on terpene extraction, the researchers discuss the various aspects of their method and provide the details of materials and equipment used. They conclude that their method "covers a large cross-section of commonly detected cannabis volatiles, is validated for a large proportion of compounds it covers, and offers significant improvement in terms of sample preparation and sample throughput over previously published studies."
Posted on July 21, 2020
By LabLynx
Journal articles
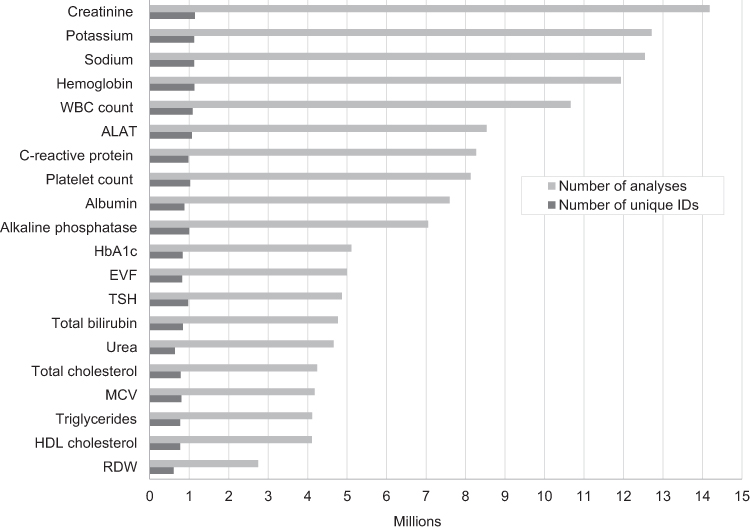
In this data resource review, researchers at several Danish hospitals discuss how the country puts two laboratory information systems (LISs) that collect routine biomarker data to use, as well as how it can be accessed for research. The researchers explain how data is collected into the LISs, how data quality is managed, and how it is used, providing several real-world examples. They then discuss the strengths and weaknesses of their data as they relate to epidemiology, as well as how the data can be accessed. They emphasize "that access to data on routine biomarkers expands the detailed biological and clinical information available on patients in the Danish healthcare system," while the "full potential is enabled through linkage to other Danish healthcare registries."
Posted on July 14, 2020
By LabLynx
Journal articles
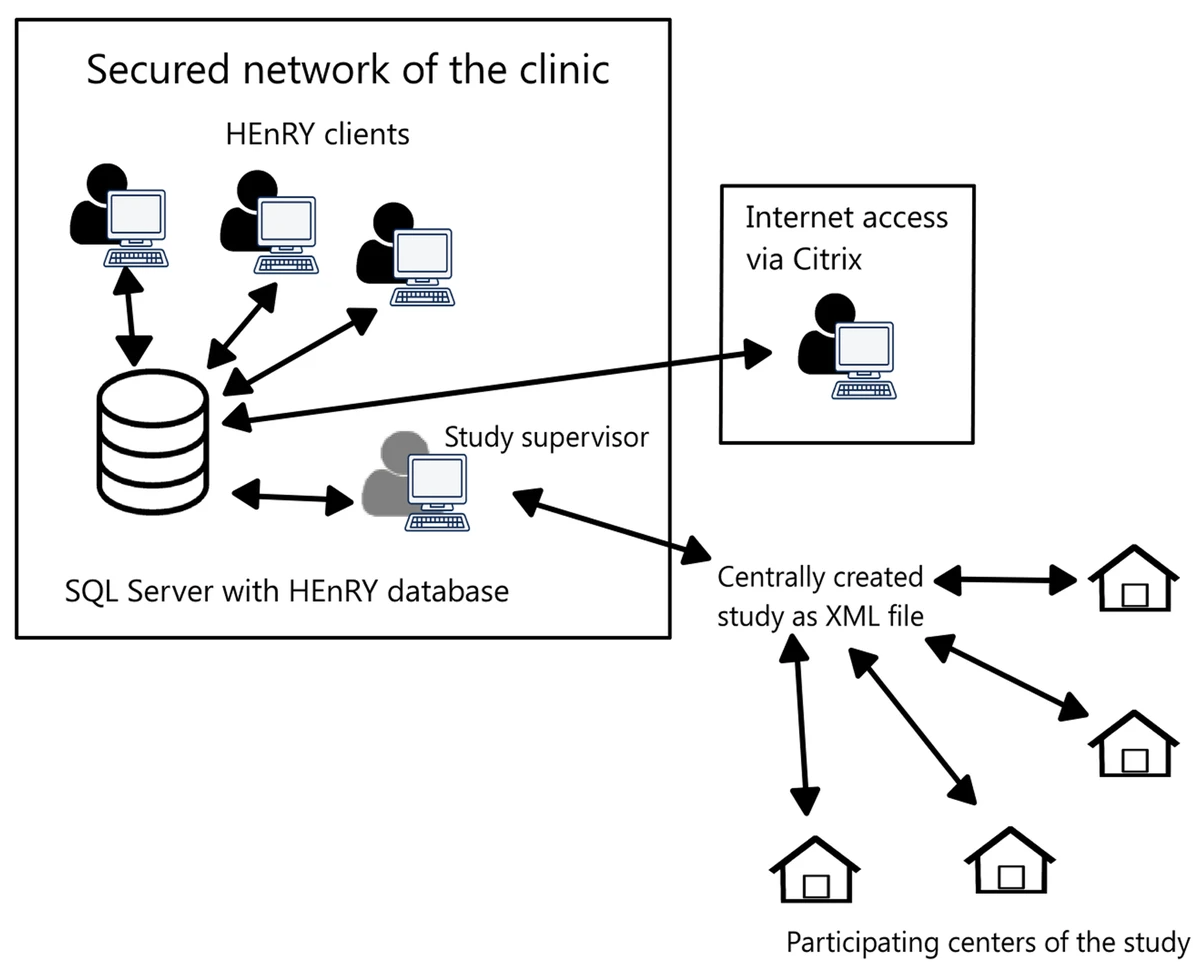
Effective biobanking of biospecimens for multicenter studies today requires more than spreadsheets and paper documents; a software system capable of improving workflows and sharing while keeping critical personal information deidentified is critical. Both commercial off-the-shelf (COTS) and open-source biobanking laboratory information management systems (LIMS) are available, but, as the University of Cologne found out, those options may be too complex to implement or cost-intensive for multicenter research networks. The university took matters into their own hands and developed the HIV Engaged Research Technology (HEnRY) LIMS, which has since expanded into a broader, open-source biobanking solution that can be applied to contexts beyond HIV research. This 2020 paper discusses the LIMS and its development and application, concluding that it offers "immense potential for emerging research groups, especially in the setting of limited resources and/or complex multicenter studies."
Posted on July 10, 2020
By LabLynx
Journal articles
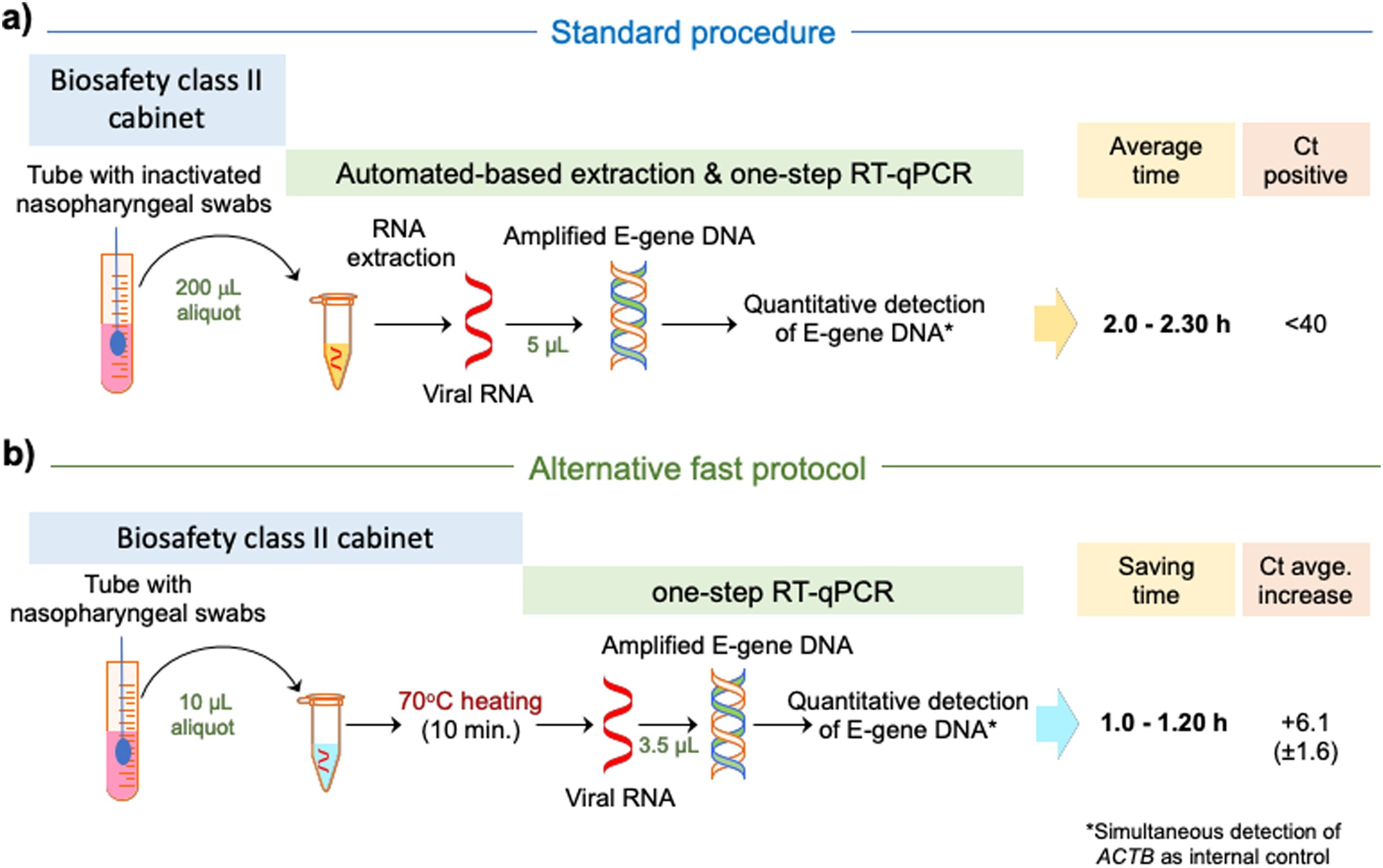
This "short communications" paper published in the
International Journal of Infectious Diseases describes the results of simplifying reverse transcription and real-time quantitative PCR (RT-qPCR) analyses for SARS-CoV-2 by skipping the RNA extraction step and instead testing three different types of direct heating of the nasopharyngeal swab. Among the three methods tested—heating directly without additives, heating with a formamide-EDTA buffer, and heating with an RNAsnap buffer—the direct heating method "provided the best results, which were highly consistent with the SARS-CoV-2 infection diagnosis based on standard RNA extraction," while also processing nearly half the time. The authors warn, however, that "choice of RT-qPCR kits might have an impact on the sensitivity of the Direct protocol" when trying to replicate their results.
Posted on July 6, 2020
By LabLynx
Journal articles

When it comes to analyzing the effects on humans of chemicals and other substances that make their way into the environment, "large, complex data sets" are often required. However, those data sets are often disparate and inconsistent. This has made epidemiological studies of air pollution and other types of environmental contamination difficult and limited in effectiveness. This problem has compounded in the age of big data. Comess
et al. address the challenges associated with today's environmental public health data in this recent paper published in
Frontiers in Artificial Intelligence and discuss how augmented intelligence, artificial intelligence (AI), and machine learning can enhance understanding of that data. However, they note, additional work must be put into improving not only analysis but also into how data is collected and shared, how researchers are trained in scientific computing and data science, and how familiarity with the benefits of AI and machine learning must be expanded. They conclude their paper with a table of five distinct challenges and their recommended way to address them so as to "create an environment that will foster the data revolution" in environmental public health and beyond.
Posted on June 23, 2020
By LabLynx
Journal articles
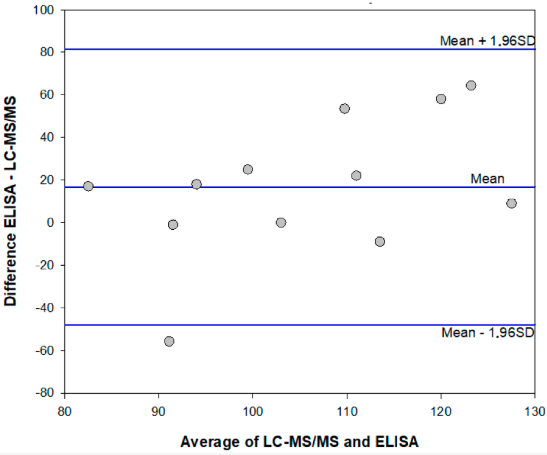
In this 2020 paper published in the journal
Toxins, Di Nardo
et al. of the University of Turin present their findings from adapting an existing enzyme immunoassay to cannabis testing for purposes of more accurately detecting mycotoxins, including aflatoxins such as aflatoxin B1 (AFB1). Citing benefits such as fewer steps, more cost-efficient equipment, and fewer training level demands, the authors viewed the application of enzyme immunoassay to testing cannabis for mycotoxins a worthy endeavor. Di Nardo
et al. discuss at length how they converted an immunoassay for measuring aflatoxins in eggs to one for the cannabis substrate, as well as the various challenges and caveats associated with the resulting methodology. In comparison to techniques such as ultra-high performance liquid chromatography coupled to high resolution tandem mass spectrometry, the authors concluded that enzyme immunoassay more readily allows for "wide applications in low resource settings and for the affordable monitoring of the safety of cannabis products, including those used recreationally and as a food supplement."
Posted on June 16, 2020
By LabLynx
Journal articles
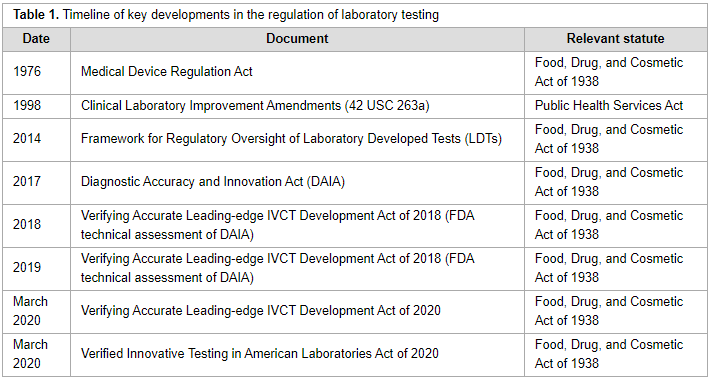
What is the current status of laboratory developed tests (LDTs) and the U.S. Food and Drug Administration's (FDA) approach to them? What are the arguments for and against their regulation? In this 2020 review article published in
Practical Laboratory Medicine, University of Washington's Eric Konnick covers the current U.S. regulatory environment for LDTs, as well as recent developments since the October 2014 FDA release of LDT draft guidance. Konnick then provides extensive commentary on the scope of LDT regulation in the U.S., including perceived risks to patients, LDT accuracy and equivalency, and how regulatory certainty fits into LDT innovation. He closes with recommendations for approaching future LDT regulation, concluding that "identifying tools that can be leveraged to improve laboratory test quality may offer many benefits that do not necessarily require a burdensome regulatory framework."
Posted on June 9, 2020
By LabLynx
Journal articles
In this 2020 article published in
Frontiers in Cell and Developmental Biology, D'Cruz
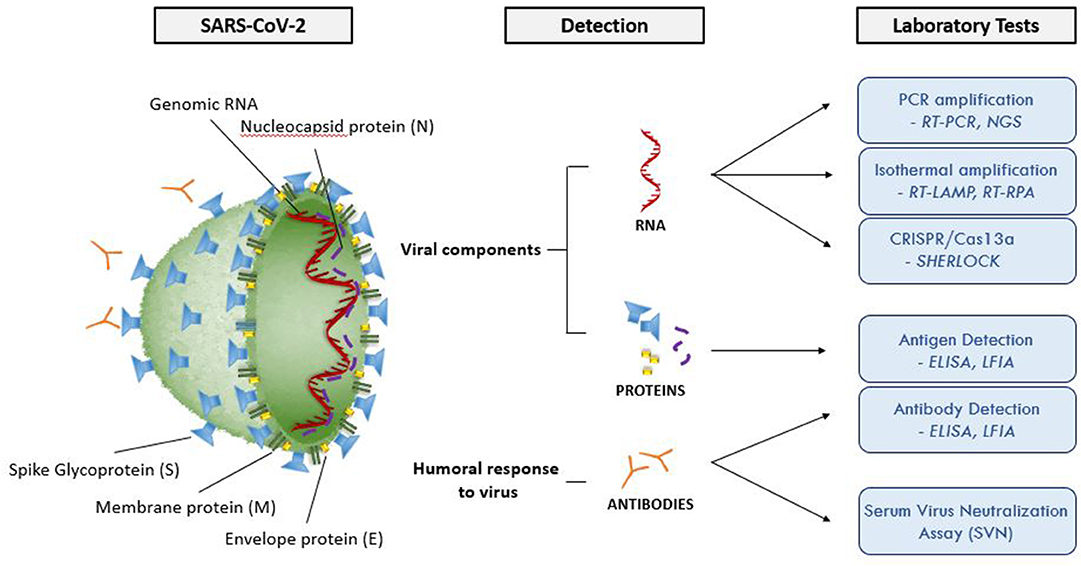
et al. provide an overview of the current laboratory methods available to test for coronaviruses, with a focus on SARS-CoV-2, the virus responsible for COVID-19. After providing an introduction to COVID-19 and its virus, the authors discuss the most common methods—such as qRT-PCR, ELISA, and LFI—as well as emerging diagnostic methods involving isothermal nucleic acid amplification, CRISPR, and NGS. They conclude with several visuals comparing the methods and when they are used, and they emphasize the importance of these test methods (as well as laboratory preparedness) in addressing rapidly evolving viral infection scenarios.
Posted on June 2, 2020
By LabLynx
Journal articles
In this 2020 article published in
Journal of Pathology Informatics, Durant
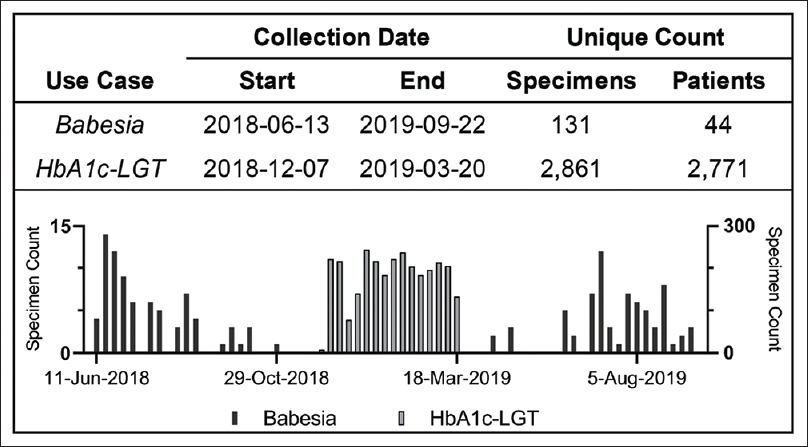
et al. of the Yale New Haven Health system present the results of their attempt to automate the identification and notification of laboratory biospecimens for biomedical researchers. Noting biospecimens' value to basic, translational, and clinical research, the authors wanted to get around the "technical, logistic, regulatory, and ethical challenges" of accessing biospecimen data. They developed Prism, a tool built on open-source technology for efficiently identifying and notifying investigators of biospecimen availability in real time. The authors present details of two use cases and conclude that their "solution is highly scalable to meet the needs of even large academic centers and reference laboratories," while also guaranteeing that the virtualization of the associated workflow "within a microservices environment does not introduce a performance penalty."
Posted on May 26, 2020
By LabLynx
Journal articles
In this 2020 journal article published in
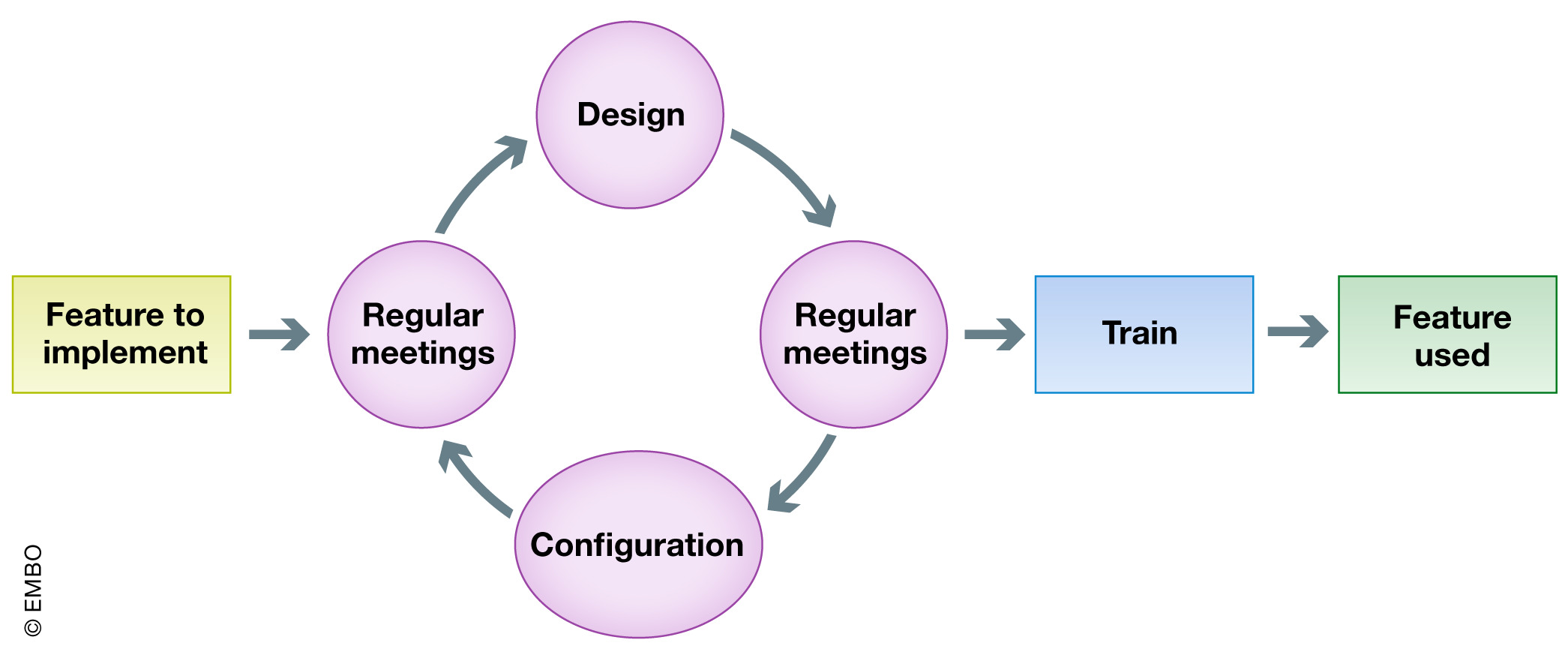
EMBO Reports, Nicolas Argento of École Polytechnique Fédérale de Lausanne (EPFL) reviews the step-by-step approach EPFL took in identifying, piloting, configuring, and implementing a combination electronic laboratory notebook (ELN)–laboratory information management system (LIMS) for member laboratories affiliated with the institution. In particular, Argento highlights the value of "surrounding yourself with the right people and the right software at the right time" when going through the various deployment steps. After providing an introduction, the author goes through the various five phases of the project they undertook, while also highlighting the value of stakeholders and other critical staff. Argento closes with a brief highlight of the "laboratory data manager" as being a role that should not be overlooked, then he concludes that while the EPFL's institutional ELN-LIMS platform meets its goals, additional data management and system adoption challenges remain.
Posted on May 18, 2020
By Shawn Douglas
Journal articles

This April 2020 whitepaper authored by Bay
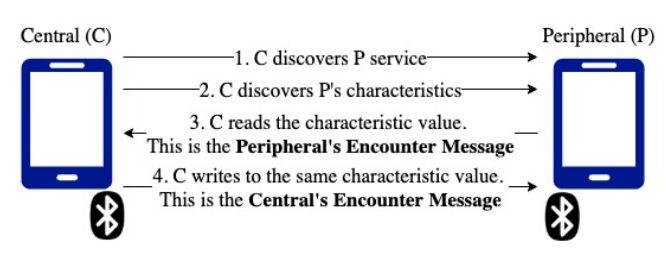
et al. of Singapore's Government Technology Agency presents BlueTrace, a privacy-preserving protocol that underpins Singapore's nationally deployed Bluetooth-based contact tracing system TraceTogether. The white paper describes the purpose of BlueTrace as addressing the key limitation of manual contact tracing: "an infected person can only report contacts they are acquainted with and remember having met." They describe the protocol's design considerations and implementation challenges, as well as how it meets interoperability requirements of health authorities considering its use. Finally they address potential security concerns surrounding the tech. They conclude that while not a stand-alone solution to solving contract tracing issues during a pandemic, Bluetooth-based automated tracing instances can still "ultimately coexist and support the pandemic response plans and processes of the public health authorities guiding us through" a pandemic. The authors also make available OpenTrace, an open-source reference implementation of BlueTrace, on GitHub.
Posted on May 12, 2020
By Shawn Douglas
Journal articles
In this 2019 paper published in
AIMS Public Health, Bhattacharya
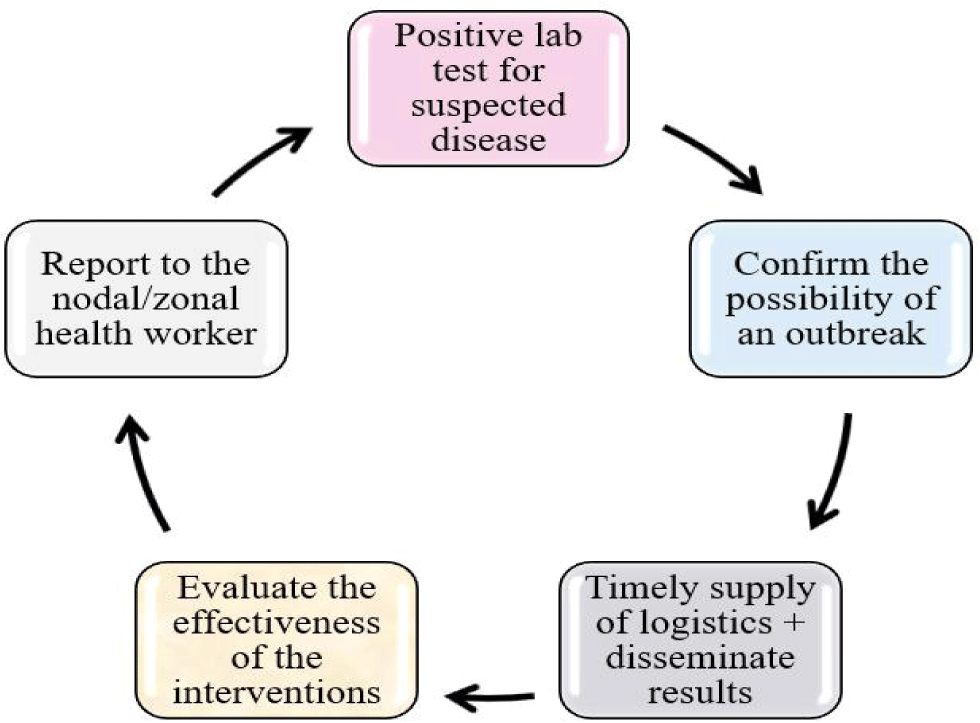
et al. argue for the benefits of using blockchain technology in public health surveillance. Noting the transparency and trust aspects of the technology, as well as the benefits of " data security and privacy, cost-effectiveness, and verifiability of data," the authors discuss four primary ways blockchain can be applied to healthcare activities, including community disease surveillance. "Blockchain could help independent organizations to manage data more efficiently during pandemics," they note, particularly in unison with machine learning and AI techniques. They conclude that blockchain-based disease surveillance "can be more effective and rapid than traditional surveillance in terms of coverage, durability, consensus, selective privacy, uniqueness, and timing."
Posted on May 5, 2020
By Shawn Douglas
Journal articles
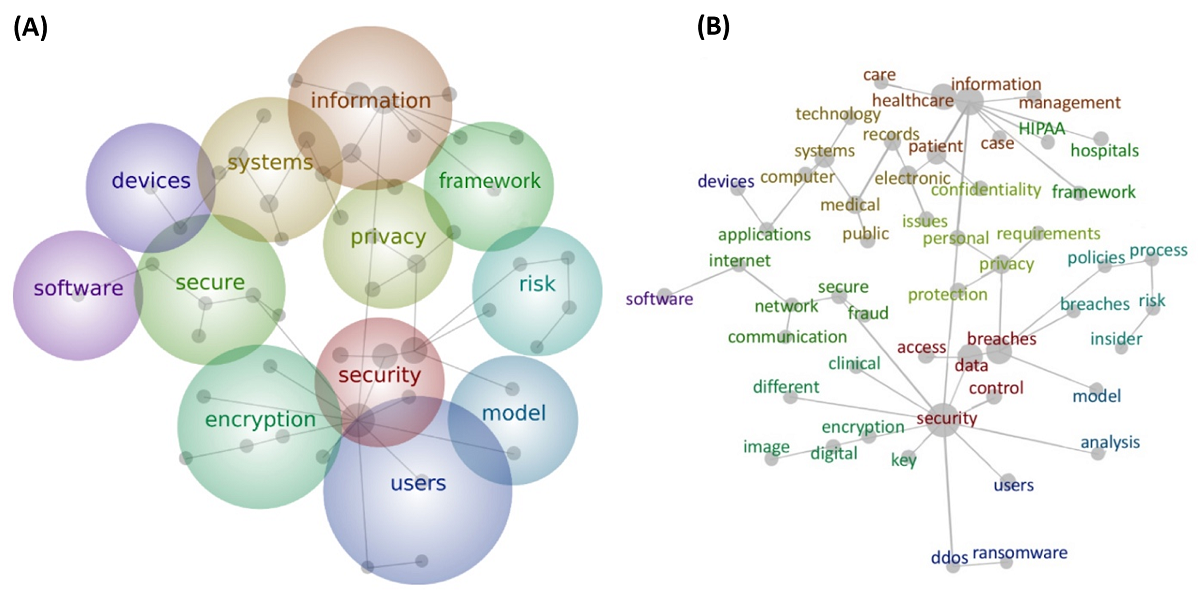
Cybersecurity remains a vital consideration for entities within the healthcare setting, with academic researchers publishing on the topic frequently. But since computer security in healthcare first started getting researched in the mid-1980s, how have topical focuses shifted? What areas, if any, are underserved by current research? Jalali
et al. attempt to answer those and other questions in this 2019 paper published in
Journal of Medical Internet Research. Using bibliometric analysis as their tool, the authors conducted chronological analysis, domain clustering analysis, and text analysis on 472 English-language articles ranging from 1985 to September 2017. The conclude that "[d]espite the increase in research and attention to cybersecurity, there are persistent shortcomings in the research on cybersecurity," including implications that emerging cyber threats, physical security methodologies, and several human elements of cybersecurity have not been sufficiently captured in research. They worry that "a school of thought that knowledge, especially specific strategies and tactics, should not be shared openly," further suppresses research growth and utility.
Posted on April 27, 2020
By Shawn Douglas
Journal articles

This week we turn back the clock a couple of years to 2018, when Fairchild
et al.published in the journal
Frontiers in Public Health their analysis of the challenges of epidemiological data reporting. With the march of COVID-19 today, their analysis and advice holds even more relevant today. In their paper, Fairchild
et al. first introduced the state of epidemiological data reporting, particularly on the internet. They then discussed the three challenges that affect such reporting: user interface, data format, and data reporting issues. They concluded with nine clear best practices that should be followed when managing epidemiological data for release to the public. They imagined a scenario of a standardized platform adopted worldwide, "where global data could be easily collected without the challenges we currently face. This would in turn streamline epidemiological and public health analysis, modeling, and informatics, resulting in better public health decision-making capabilities."
Posted on April 20, 2020
By Shawn Douglas
Journal articles
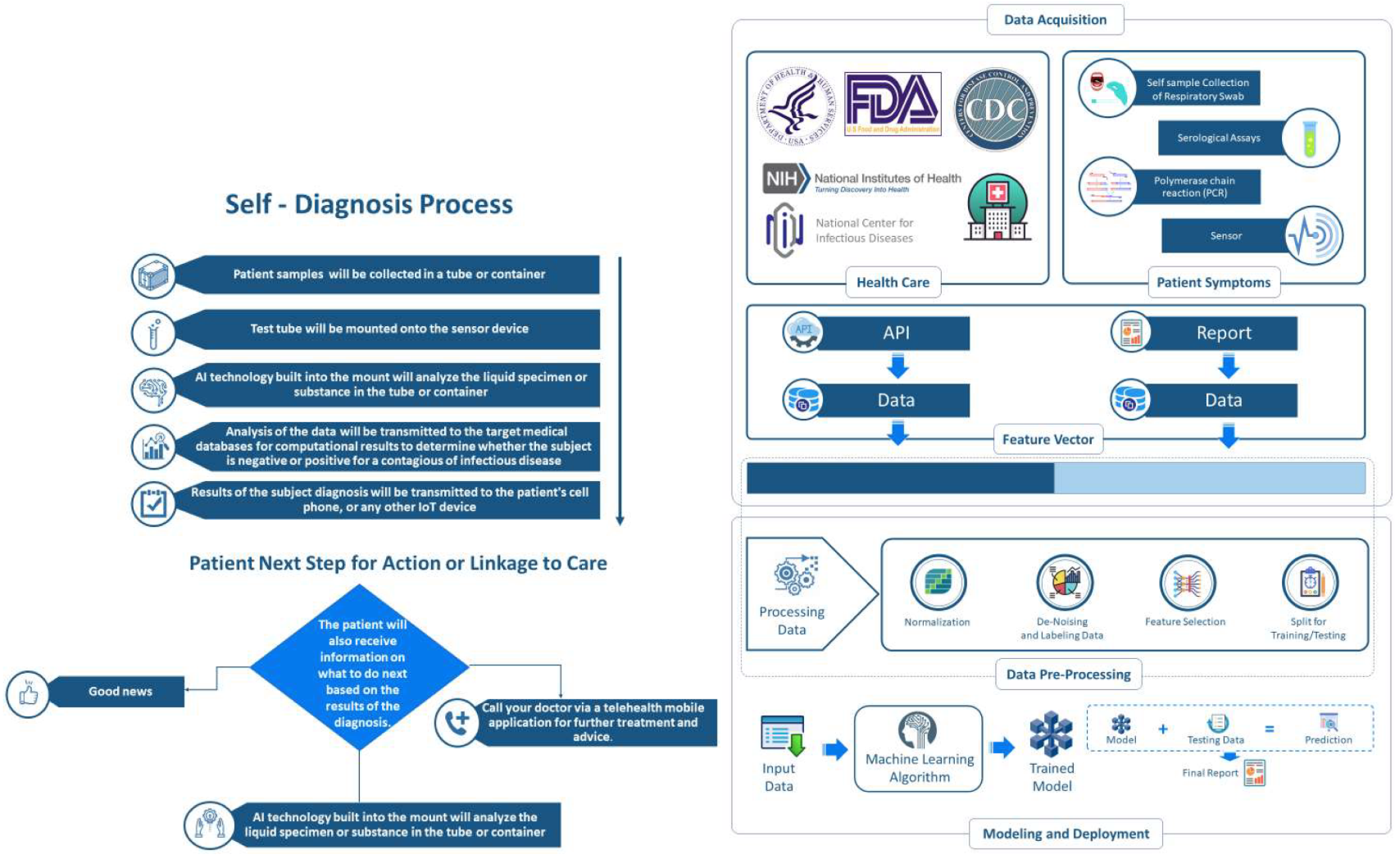
In this brief editorial in the journal
Diagnostics, Mashamba-Thompson
et al. state their case for a potentially ideal solution to COVID-19 testing in resource-poor communities: using mobile health (mHealth) solutions in conjunction with a blockchain and AI-driven data acquisition and transfer system. They add that the "AI component of this technology enables powerful data collection (patient information, geographic location of the patient, and test results), security, analysis, and curation of disparate and clinical data sets from federated blockchain platforms to derive triangulated data at very high degrees of confidence and speed. " Unfortunately, what isn't clear in their argument is how the self-testing step would actually work, let alone how the system would realistically be implemented in resource-poor settings. One could argue, however, despite the relatively few details, the sharing of ideas on how to address COVID-19 testing in various environments still has value.
Posted on April 13, 2020
By Shawn Douglas
Journal articles
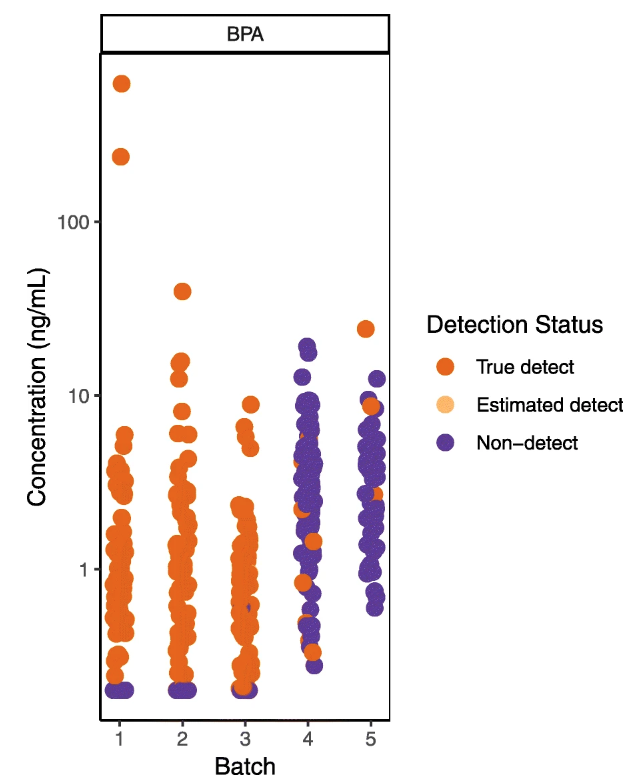
In this 2019 paper published in
Environmental Health, Udesky
et al. share their organization's guidance towards addressing quality in environmental exposure testing and data. In particular, the authors note the importance of reporting quality control data along with chemical measurements in published research, stating that its lacking leaves "readers uncertain about the level of confidence in the reported data." Their objective was to provide full guidance on the implementation, interpretation, and reporting of QC data. After explaining their approach to study design, study implementation, and data interpretation, the authors conclude that their guidance, visualizations, and supplementary content provide "a useful set of tools for getting the best information from valuable environmental exposure datasets, and enabling valid comparison and synthesis of exposure data across studies."
Posted on April 6, 2020
By Shawn Douglas
Journal articles
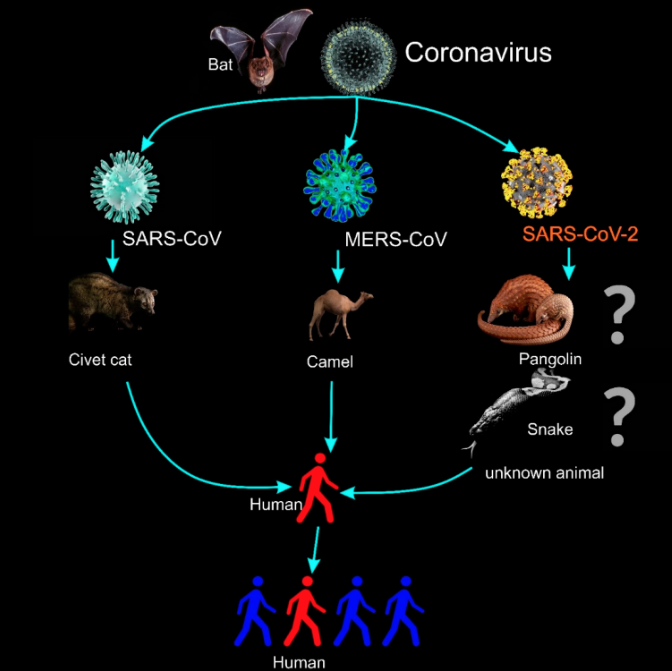
In this March 2020 journal article published in the journal
Micromachines, Nguyen, Bang, and Wolff give their professional take on point-of-care diagnostics for combating the COVID-19 epidemic. In particular, the authors discuss the loop-mediated isothermal amplification (LAMP) nucleic acid amplification technique and its potential to make COVID-19 testing faster, more portable, more flexible, more stable, and more sensitive. After discussing the then-recent events (information is changing rapidly), the authors compare real-time reverse transcription polymerase chain reaction (rRT-PCR) with LAMP methods and discuss other important factors affecting society because of the pandemic.
Posted on March 30, 2020
By Shawn Douglas
Journal articles
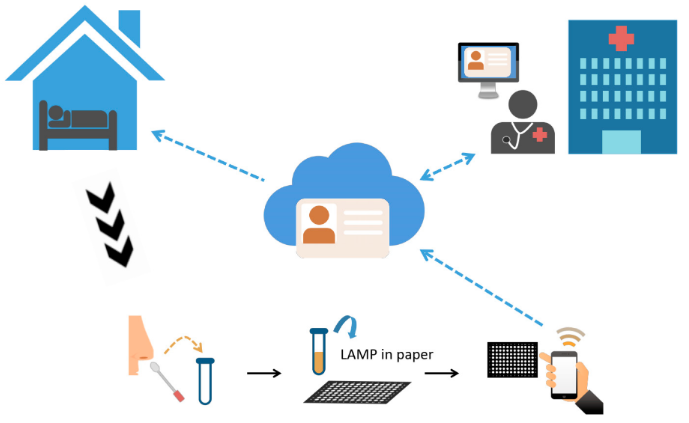
In this brief editorial article published in the journal
Diagnostics, Tiawanese researchers Yang
et al. provide their insights into a more convenient at-home point-of-care (POC) testing method for the COVID-19 illness. Noting the urgency of the associated pandemic and the various costs associated with traditional testing methodologies such as lateral flow immunoassays and molecular-based assays, the authors suggest the blending of several tools: a paper-based result and common mobile devices. A patient at home could take a nasal swab and use a POC device to return a colorimetric result, which could then be captured by mobile phone and rapidly sent to a clinician for analysis. They conclude that such a method could "provide new insights into designing POC COVID-19 diagnostics and ultimately improve the health care system to combat this and similar diseases."
 In this 2020 short report from Dixon et al., lessons learned from attempts to expand data quality measures in the open-source tool Observational Health Data Science and Informatics (OHDSI) are presented. Noting a lack of data quality assessment and improvement mechanisms in health information systems in general, the researchers sought to improve OHDSI for public health surveillance use cases. After explaining the practical uses of OHDSI, the authors state their case for why measuring completeness, timeliness, and information entropy within OHDSI would be useful. Though not getting approval to add timeliness to the system, they conclude that high value remains "in adapting existing infrastructure and tools to support expanded use cases rather than to just create independent tools for use by a niche group."
In this 2020 short report from Dixon et al., lessons learned from attempts to expand data quality measures in the open-source tool Observational Health Data Science and Informatics (OHDSI) are presented. Noting a lack of data quality assessment and improvement mechanisms in health information systems in general, the researchers sought to improve OHDSI for public health surveillance use cases. After explaining the practical uses of OHDSI, the authors state their case for why measuring completeness, timeliness, and information entropy within OHDSI would be useful. Though not getting approval to add timeliness to the system, they conclude that high value remains "in adapting existing infrastructure and tools to support expanded use cases rather than to just create independent tools for use by a niche group."
 This February 2020 paper published in BMC Medical Informatics and Decision Making examines the state of health information exchange in Québec and other parts of Canada and how its application to laboratory medicine might be improved. In particular, laboratory information exchange (LIE) systems that "improve the reliability of the laboratory testing process" and integrate "with other clinical information systems (CISs) physicians use in hospitals" are examined in this work. Surveying hospital-based specialist physicians, Raymond et al. paint a picture of how varying clinical information management solutions are used, what functionality is being used and not used, and how physicians view the potential benefits of the clinical systems they use. They conclude that there is very much a "complementary nature" between systems and that "system designers should take a step back to imagine a way to design systems as part of an interconnected network of features."
This February 2020 paper published in BMC Medical Informatics and Decision Making examines the state of health information exchange in Québec and other parts of Canada and how its application to laboratory medicine might be improved. In particular, laboratory information exchange (LIE) systems that "improve the reliability of the laboratory testing process" and integrate "with other clinical information systems (CISs) physicians use in hospitals" are examined in this work. Surveying hospital-based specialist physicians, Raymond et al. paint a picture of how varying clinical information management solutions are used, what functionality is being used and not used, and how physicians view the potential benefits of the clinical systems they use. They conclude that there is very much a "complementary nature" between systems and that "system designers should take a step back to imagine a way to design systems as part of an interconnected network of features."
 In this 2020 paper published in the journal Matabolites, Krill et al. of Australia's AgriBio present their findings in an attempt to reduce runtimes and extraction complexity associated with quantifying terpenes in cannabis biomass. Noting the evolving needs of high-throughput cannabis breeding programs, the researchers present a method "based on a simple hexane extract from 40 mg of biomass, with 50 μg/mL dodecane as internal standard, and a gradient of less than 30 minutes." After presenting current background on terpene extraction, the researchers discuss the various aspects of their method and provide the details of materials and equipment used. They conclude that their method "covers a large cross-section of commonly detected cannabis volatiles, is validated for a large proportion of compounds it covers, and offers significant improvement in terms of sample preparation and sample throughput over previously published studies."
In this 2020 paper published in the journal Matabolites, Krill et al. of Australia's AgriBio present their findings in an attempt to reduce runtimes and extraction complexity associated with quantifying terpenes in cannabis biomass. Noting the evolving needs of high-throughput cannabis breeding programs, the researchers present a method "based on a simple hexane extract from 40 mg of biomass, with 50 μg/mL dodecane as internal standard, and a gradient of less than 30 minutes." After presenting current background on terpene extraction, the researchers discuss the various aspects of their method and provide the details of materials and equipment used. They conclude that their method "covers a large cross-section of commonly detected cannabis volatiles, is validated for a large proportion of compounds it covers, and offers significant improvement in terms of sample preparation and sample throughput over previously published studies."
 In this data resource review, researchers at several Danish hospitals discuss how the country puts two laboratory information systems (LISs) that collect routine biomarker data to use, as well as how it can be accessed for research. The researchers explain how data is collected into the LISs, how data quality is managed, and how it is used, providing several real-world examples. They then discuss the strengths and weaknesses of their data as they relate to epidemiology, as well as how the data can be accessed. They emphasize "that access to data on routine biomarkers expands the detailed biological and clinical information available on patients in the Danish healthcare system," while the "full potential is enabled through linkage to other Danish healthcare registries."
In this data resource review, researchers at several Danish hospitals discuss how the country puts two laboratory information systems (LISs) that collect routine biomarker data to use, as well as how it can be accessed for research. The researchers explain how data is collected into the LISs, how data quality is managed, and how it is used, providing several real-world examples. They then discuss the strengths and weaknesses of their data as they relate to epidemiology, as well as how the data can be accessed. They emphasize "that access to data on routine biomarkers expands the detailed biological and clinical information available on patients in the Danish healthcare system," while the "full potential is enabled through linkage to other Danish healthcare registries."
 Effective biobanking of biospecimens for multicenter studies today requires more than spreadsheets and paper documents; a software system capable of improving workflows and sharing while keeping critical personal information deidentified is critical. Both commercial off-the-shelf (COTS) and open-source biobanking laboratory information management systems (LIMS) are available, but, as the University of Cologne found out, those options may be too complex to implement or cost-intensive for multicenter research networks. The university took matters into their own hands and developed the HIV Engaged Research Technology (HEnRY) LIMS, which has since expanded into a broader, open-source biobanking solution that can be applied to contexts beyond HIV research. This 2020 paper discusses the LIMS and its development and application, concluding that it offers "immense potential for emerging research groups, especially in the setting of limited resources and/or complex multicenter studies."
Effective biobanking of biospecimens for multicenter studies today requires more than spreadsheets and paper documents; a software system capable of improving workflows and sharing while keeping critical personal information deidentified is critical. Both commercial off-the-shelf (COTS) and open-source biobanking laboratory information management systems (LIMS) are available, but, as the University of Cologne found out, those options may be too complex to implement or cost-intensive for multicenter research networks. The university took matters into their own hands and developed the HIV Engaged Research Technology (HEnRY) LIMS, which has since expanded into a broader, open-source biobanking solution that can be applied to contexts beyond HIV research. This 2020 paper discusses the LIMS and its development and application, concluding that it offers "immense potential for emerging research groups, especially in the setting of limited resources and/or complex multicenter studies."
 This "short communications" paper published in the International Journal of Infectious Diseases describes the results of simplifying reverse transcription and real-time quantitative PCR (RT-qPCR) analyses for SARS-CoV-2 by skipping the RNA extraction step and instead testing three different types of direct heating of the nasopharyngeal swab. Among the three methods tested—heating directly without additives, heating with a formamide-EDTA buffer, and heating with an RNAsnap buffer—the direct heating method "provided the best results, which were highly consistent with the SARS-CoV-2 infection diagnosis based on standard RNA extraction," while also processing nearly half the time. The authors warn, however, that "choice of RT-qPCR kits might have an impact on the sensitivity of the Direct protocol" when trying to replicate their results.
This "short communications" paper published in the International Journal of Infectious Diseases describes the results of simplifying reverse transcription and real-time quantitative PCR (RT-qPCR) analyses for SARS-CoV-2 by skipping the RNA extraction step and instead testing three different types of direct heating of the nasopharyngeal swab. Among the three methods tested—heating directly without additives, heating with a formamide-EDTA buffer, and heating with an RNAsnap buffer—the direct heating method "provided the best results, which were highly consistent with the SARS-CoV-2 infection diagnosis based on standard RNA extraction," while also processing nearly half the time. The authors warn, however, that "choice of RT-qPCR kits might have an impact on the sensitivity of the Direct protocol" when trying to replicate their results.
 When it comes to analyzing the effects on humans of chemicals and other substances that make their way into the environment, "large, complex data sets" are often required. However, those data sets are often disparate and inconsistent. This has made epidemiological studies of air pollution and other types of environmental contamination difficult and limited in effectiveness. This problem has compounded in the age of big data. Comess et al. address the challenges associated with today's environmental public health data in this recent paper published in Frontiers in Artificial Intelligence and discuss how augmented intelligence, artificial intelligence (AI), and machine learning can enhance understanding of that data. However, they note, additional work must be put into improving not only analysis but also into how data is collected and shared, how researchers are trained in scientific computing and data science, and how familiarity with the benefits of AI and machine learning must be expanded. They conclude their paper with a table of five distinct challenges and their recommended way to address them so as to "create an environment that will foster the data revolution" in environmental public health and beyond.
When it comes to analyzing the effects on humans of chemicals and other substances that make their way into the environment, "large, complex data sets" are often required. However, those data sets are often disparate and inconsistent. This has made epidemiological studies of air pollution and other types of environmental contamination difficult and limited in effectiveness. This problem has compounded in the age of big data. Comess et al. address the challenges associated with today's environmental public health data in this recent paper published in Frontiers in Artificial Intelligence and discuss how augmented intelligence, artificial intelligence (AI), and machine learning can enhance understanding of that data. However, they note, additional work must be put into improving not only analysis but also into how data is collected and shared, how researchers are trained in scientific computing and data science, and how familiarity with the benefits of AI and machine learning must be expanded. They conclude their paper with a table of five distinct challenges and their recommended way to address them so as to "create an environment that will foster the data revolution" in environmental public health and beyond.
 In this 2020 paper published in the journal Toxins, Di Nardo et al. of the University of Turin present their findings from adapting an existing enzyme immunoassay to cannabis testing for purposes of more accurately detecting mycotoxins, including aflatoxins such as aflatoxin B1 (AFB1). Citing benefits such as fewer steps, more cost-efficient equipment, and fewer training level demands, the authors viewed the application of enzyme immunoassay to testing cannabis for mycotoxins a worthy endeavor. Di Nardo et al. discuss at length how they converted an immunoassay for measuring aflatoxins in eggs to one for the cannabis substrate, as well as the various challenges and caveats associated with the resulting methodology. In comparison to techniques such as ultra-high performance liquid chromatography coupled to high resolution tandem mass spectrometry, the authors concluded that enzyme immunoassay more readily allows for "wide applications in low resource settings and for the affordable monitoring of the safety of cannabis products, including those used recreationally and as a food supplement."
In this 2020 paper published in the journal Toxins, Di Nardo et al. of the University of Turin present their findings from adapting an existing enzyme immunoassay to cannabis testing for purposes of more accurately detecting mycotoxins, including aflatoxins such as aflatoxin B1 (AFB1). Citing benefits such as fewer steps, more cost-efficient equipment, and fewer training level demands, the authors viewed the application of enzyme immunoassay to testing cannabis for mycotoxins a worthy endeavor. Di Nardo et al. discuss at length how they converted an immunoassay for measuring aflatoxins in eggs to one for the cannabis substrate, as well as the various challenges and caveats associated with the resulting methodology. In comparison to techniques such as ultra-high performance liquid chromatography coupled to high resolution tandem mass spectrometry, the authors concluded that enzyme immunoassay more readily allows for "wide applications in low resource settings and for the affordable monitoring of the safety of cannabis products, including those used recreationally and as a food supplement."
 What is the current status of laboratory developed tests (LDTs) and the U.S. Food and Drug Administration's (FDA) approach to them? What are the arguments for and against their regulation? In this 2020 review article published in Practical Laboratory Medicine, University of Washington's Eric Konnick covers the current U.S. regulatory environment for LDTs, as well as recent developments since the October 2014 FDA release of LDT draft guidance. Konnick then provides extensive commentary on the scope of LDT regulation in the U.S., including perceived risks to patients, LDT accuracy and equivalency, and how regulatory certainty fits into LDT innovation. He closes with recommendations for approaching future LDT regulation, concluding that "identifying tools that can be leveraged to improve laboratory test quality may offer many benefits that do not necessarily require a burdensome regulatory framework."
What is the current status of laboratory developed tests (LDTs) and the U.S. Food and Drug Administration's (FDA) approach to them? What are the arguments for and against their regulation? In this 2020 review article published in Practical Laboratory Medicine, University of Washington's Eric Konnick covers the current U.S. regulatory environment for LDTs, as well as recent developments since the October 2014 FDA release of LDT draft guidance. Konnick then provides extensive commentary on the scope of LDT regulation in the U.S., including perceived risks to patients, LDT accuracy and equivalency, and how regulatory certainty fits into LDT innovation. He closes with recommendations for approaching future LDT regulation, concluding that "identifying tools that can be leveraged to improve laboratory test quality may offer many benefits that do not necessarily require a burdensome regulatory framework."
 In this 2020 article published in Frontiers in Cell and Developmental Biology, D'Cruz et al. provide an overview of the current laboratory methods available to test for coronaviruses, with a focus on SARS-CoV-2, the virus responsible for COVID-19. After providing an introduction to COVID-19 and its virus, the authors discuss the most common methods—such as qRT-PCR, ELISA, and LFI—as well as emerging diagnostic methods involving isothermal nucleic acid amplification, CRISPR, and NGS. They conclude with several visuals comparing the methods and when they are used, and they emphasize the importance of these test methods (as well as laboratory preparedness) in addressing rapidly evolving viral infection scenarios.
In this 2020 article published in Frontiers in Cell and Developmental Biology, D'Cruz et al. provide an overview of the current laboratory methods available to test for coronaviruses, with a focus on SARS-CoV-2, the virus responsible for COVID-19. After providing an introduction to COVID-19 and its virus, the authors discuss the most common methods—such as qRT-PCR, ELISA, and LFI—as well as emerging diagnostic methods involving isothermal nucleic acid amplification, CRISPR, and NGS. They conclude with several visuals comparing the methods and when they are used, and they emphasize the importance of these test methods (as well as laboratory preparedness) in addressing rapidly evolving viral infection scenarios.
 In this 2020 article published in Journal of Pathology Informatics, Durant et al. of the Yale New Haven Health system present the results of their attempt to automate the identification and notification of laboratory biospecimens for biomedical researchers. Noting biospecimens' value to basic, translational, and clinical research, the authors wanted to get around the "technical, logistic, regulatory, and ethical challenges" of accessing biospecimen data. They developed Prism, a tool built on open-source technology for efficiently identifying and notifying investigators of biospecimen availability in real time. The authors present details of two use cases and conclude that their "solution is highly scalable to meet the needs of even large academic centers and reference laboratories," while also guaranteeing that the virtualization of the associated workflow "within a microservices environment does not introduce a performance penalty."
In this 2020 article published in Journal of Pathology Informatics, Durant et al. of the Yale New Haven Health system present the results of their attempt to automate the identification and notification of laboratory biospecimens for biomedical researchers. Noting biospecimens' value to basic, translational, and clinical research, the authors wanted to get around the "technical, logistic, regulatory, and ethical challenges" of accessing biospecimen data. They developed Prism, a tool built on open-source technology for efficiently identifying and notifying investigators of biospecimen availability in real time. The authors present details of two use cases and conclude that their "solution is highly scalable to meet the needs of even large academic centers and reference laboratories," while also guaranteeing that the virtualization of the associated workflow "within a microservices environment does not introduce a performance penalty."
 In this 2020 journal article published in EMBO Reports, Nicolas Argento of École Polytechnique Fédérale de Lausanne (EPFL) reviews the step-by-step approach EPFL took in identifying, piloting, configuring, and implementing a combination electronic laboratory notebook (ELN)–laboratory information management system (LIMS) for member laboratories affiliated with the institution. In particular, Argento highlights the value of "surrounding yourself with the right people and the right software at the right time" when going through the various deployment steps. After providing an introduction, the author goes through the various five phases of the project they undertook, while also highlighting the value of stakeholders and other critical staff. Argento closes with a brief highlight of the "laboratory data manager" as being a role that should not be overlooked, then he concludes that while the EPFL's institutional ELN-LIMS platform meets its goals, additional data management and system adoption challenges remain.
In this 2020 journal article published in EMBO Reports, Nicolas Argento of École Polytechnique Fédérale de Lausanne (EPFL) reviews the step-by-step approach EPFL took in identifying, piloting, configuring, and implementing a combination electronic laboratory notebook (ELN)–laboratory information management system (LIMS) for member laboratories affiliated with the institution. In particular, Argento highlights the value of "surrounding yourself with the right people and the right software at the right time" when going through the various deployment steps. After providing an introduction, the author goes through the various five phases of the project they undertook, while also highlighting the value of stakeholders and other critical staff. Argento closes with a brief highlight of the "laboratory data manager" as being a role that should not be overlooked, then he concludes that while the EPFL's institutional ELN-LIMS platform meets its goals, additional data management and system adoption challenges remain.
 This April 2020 whitepaper authored by Bay et al. of Singapore's Government Technology Agency presents BlueTrace, a privacy-preserving protocol that underpins Singapore's nationally deployed Bluetooth-based contact tracing system TraceTogether. The white paper describes the purpose of BlueTrace as addressing the key limitation of manual contact tracing: "an infected person can only report contacts they are acquainted with and remember having met." They describe the protocol's design considerations and implementation challenges, as well as how it meets interoperability requirements of health authorities considering its use. Finally they address potential security concerns surrounding the tech. They conclude that while not a stand-alone solution to solving contract tracing issues during a pandemic, Bluetooth-based automated tracing instances can still "ultimately coexist and support the pandemic response plans and processes of the public health authorities guiding us through" a pandemic. The authors also make available OpenTrace, an open-source reference implementation of BlueTrace, on GitHub.
This April 2020 whitepaper authored by Bay et al. of Singapore's Government Technology Agency presents BlueTrace, a privacy-preserving protocol that underpins Singapore's nationally deployed Bluetooth-based contact tracing system TraceTogether. The white paper describes the purpose of BlueTrace as addressing the key limitation of manual contact tracing: "an infected person can only report contacts they are acquainted with and remember having met." They describe the protocol's design considerations and implementation challenges, as well as how it meets interoperability requirements of health authorities considering its use. Finally they address potential security concerns surrounding the tech. They conclude that while not a stand-alone solution to solving contract tracing issues during a pandemic, Bluetooth-based automated tracing instances can still "ultimately coexist and support the pandemic response plans and processes of the public health authorities guiding us through" a pandemic. The authors also make available OpenTrace, an open-source reference implementation of BlueTrace, on GitHub.
 In this 2019 paper published in AIMS Public Health, Bhattacharya et al. argue for the benefits of using blockchain technology in public health surveillance. Noting the transparency and trust aspects of the technology, as well as the benefits of " data security and privacy, cost-effectiveness, and verifiability of data," the authors discuss four primary ways blockchain can be applied to healthcare activities, including community disease surveillance. "Blockchain could help independent organizations to manage data more efficiently during pandemics," they note, particularly in unison with machine learning and AI techniques. They conclude that blockchain-based disease surveillance "can be more effective and rapid than traditional surveillance in terms of coverage, durability, consensus, selective privacy, uniqueness, and timing."
In this 2019 paper published in AIMS Public Health, Bhattacharya et al. argue for the benefits of using blockchain technology in public health surveillance. Noting the transparency and trust aspects of the technology, as well as the benefits of " data security and privacy, cost-effectiveness, and verifiability of data," the authors discuss four primary ways blockchain can be applied to healthcare activities, including community disease surveillance. "Blockchain could help independent organizations to manage data more efficiently during pandemics," they note, particularly in unison with machine learning and AI techniques. They conclude that blockchain-based disease surveillance "can be more effective and rapid than traditional surveillance in terms of coverage, durability, consensus, selective privacy, uniqueness, and timing."
 Cybersecurity remains a vital consideration for entities within the healthcare setting, with academic researchers publishing on the topic frequently. But since computer security in healthcare first started getting researched in the mid-1980s, how have topical focuses shifted? What areas, if any, are underserved by current research? Jalali et al. attempt to answer those and other questions in this 2019 paper published in Journal of Medical Internet Research. Using bibliometric analysis as their tool, the authors conducted chronological analysis, domain clustering analysis, and text analysis on 472 English-language articles ranging from 1985 to September 2017. The conclude that "[d]espite the increase in research and attention to cybersecurity, there are persistent shortcomings in the research on cybersecurity," including implications that emerging cyber threats, physical security methodologies, and several human elements of cybersecurity have not been sufficiently captured in research. They worry that "a school of thought that knowledge, especially specific strategies and tactics, should not be shared openly," further suppresses research growth and utility.
Cybersecurity remains a vital consideration for entities within the healthcare setting, with academic researchers publishing on the topic frequently. But since computer security in healthcare first started getting researched in the mid-1980s, how have topical focuses shifted? What areas, if any, are underserved by current research? Jalali et al. attempt to answer those and other questions in this 2019 paper published in Journal of Medical Internet Research. Using bibliometric analysis as their tool, the authors conducted chronological analysis, domain clustering analysis, and text analysis on 472 English-language articles ranging from 1985 to September 2017. The conclude that "[d]espite the increase in research and attention to cybersecurity, there are persistent shortcomings in the research on cybersecurity," including implications that emerging cyber threats, physical security methodologies, and several human elements of cybersecurity have not been sufficiently captured in research. They worry that "a school of thought that knowledge, especially specific strategies and tactics, should not be shared openly," further suppresses research growth and utility.
 This week we turn back the clock a couple of years to 2018, when Fairchild et al.published in the journal Frontiers in Public Health their analysis of the challenges of epidemiological data reporting. With the march of COVID-19 today, their analysis and advice holds even more relevant today. In their paper, Fairchild et al. first introduced the state of epidemiological data reporting, particularly on the internet. They then discussed the three challenges that affect such reporting: user interface, data format, and data reporting issues. They concluded with nine clear best practices that should be followed when managing epidemiological data for release to the public. They imagined a scenario of a standardized platform adopted worldwide, "where global data could be easily collected without the challenges we currently face. This would in turn streamline epidemiological and public health analysis, modeling, and informatics, resulting in better public health decision-making capabilities."
This week we turn back the clock a couple of years to 2018, when Fairchild et al.published in the journal Frontiers in Public Health their analysis of the challenges of epidemiological data reporting. With the march of COVID-19 today, their analysis and advice holds even more relevant today. In their paper, Fairchild et al. first introduced the state of epidemiological data reporting, particularly on the internet. They then discussed the three challenges that affect such reporting: user interface, data format, and data reporting issues. They concluded with nine clear best practices that should be followed when managing epidemiological data for release to the public. They imagined a scenario of a standardized platform adopted worldwide, "where global data could be easily collected without the challenges we currently face. This would in turn streamline epidemiological and public health analysis, modeling, and informatics, resulting in better public health decision-making capabilities."
 In this brief editorial in the journal Diagnostics, Mashamba-Thompson et al. state their case for a potentially ideal solution to COVID-19 testing in resource-poor communities: using mobile health (mHealth) solutions in conjunction with a blockchain and AI-driven data acquisition and transfer system. They add that the "AI component of this technology enables powerful data collection (patient information, geographic location of the patient, and test results), security, analysis, and curation of disparate and clinical data sets from federated blockchain platforms to derive triangulated data at very high degrees of confidence and speed. " Unfortunately, what isn't clear in their argument is how the self-testing step would actually work, let alone how the system would realistically be implemented in resource-poor settings. One could argue, however, despite the relatively few details, the sharing of ideas on how to address COVID-19 testing in various environments still has value.
In this brief editorial in the journal Diagnostics, Mashamba-Thompson et al. state their case for a potentially ideal solution to COVID-19 testing in resource-poor communities: using mobile health (mHealth) solutions in conjunction with a blockchain and AI-driven data acquisition and transfer system. They add that the "AI component of this technology enables powerful data collection (patient information, geographic location of the patient, and test results), security, analysis, and curation of disparate and clinical data sets from federated blockchain platforms to derive triangulated data at very high degrees of confidence and speed. " Unfortunately, what isn't clear in their argument is how the self-testing step would actually work, let alone how the system would realistically be implemented in resource-poor settings. One could argue, however, despite the relatively few details, the sharing of ideas on how to address COVID-19 testing in various environments still has value.
 In this 2019 paper published in Environmental Health, Udesky et al. share their organization's guidance towards addressing quality in environmental exposure testing and data. In particular, the authors note the importance of reporting quality control data along with chemical measurements in published research, stating that its lacking leaves "readers uncertain about the level of confidence in the reported data." Their objective was to provide full guidance on the implementation, interpretation, and reporting of QC data. After explaining their approach to study design, study implementation, and data interpretation, the authors conclude that their guidance, visualizations, and supplementary content provide "a useful set of tools for getting the best information from valuable environmental exposure datasets, and enabling valid comparison and synthesis of exposure data across studies."
In this 2019 paper published in Environmental Health, Udesky et al. share their organization's guidance towards addressing quality in environmental exposure testing and data. In particular, the authors note the importance of reporting quality control data along with chemical measurements in published research, stating that its lacking leaves "readers uncertain about the level of confidence in the reported data." Their objective was to provide full guidance on the implementation, interpretation, and reporting of QC data. After explaining their approach to study design, study implementation, and data interpretation, the authors conclude that their guidance, visualizations, and supplementary content provide "a useful set of tools for getting the best information from valuable environmental exposure datasets, and enabling valid comparison and synthesis of exposure data across studies."
 In this March 2020 journal article published in the journal Micromachines, Nguyen, Bang, and Wolff give their professional take on point-of-care diagnostics for combating the COVID-19 epidemic. In particular, the authors discuss the loop-mediated isothermal amplification (LAMP) nucleic acid amplification technique and its potential to make COVID-19 testing faster, more portable, more flexible, more stable, and more sensitive. After discussing the then-recent events (information is changing rapidly), the authors compare real-time reverse transcription polymerase chain reaction (rRT-PCR) with LAMP methods and discuss other important factors affecting society because of the pandemic.
In this March 2020 journal article published in the journal Micromachines, Nguyen, Bang, and Wolff give their professional take on point-of-care diagnostics for combating the COVID-19 epidemic. In particular, the authors discuss the loop-mediated isothermal amplification (LAMP) nucleic acid amplification technique and its potential to make COVID-19 testing faster, more portable, more flexible, more stable, and more sensitive. After discussing the then-recent events (information is changing rapidly), the authors compare real-time reverse transcription polymerase chain reaction (rRT-PCR) with LAMP methods and discuss other important factors affecting society because of the pandemic.
 In this brief editorial article published in the journal Diagnostics, Tiawanese researchers Yang et al. provide their insights into a more convenient at-home point-of-care (POC) testing method for the COVID-19 illness. Noting the urgency of the associated pandemic and the various costs associated with traditional testing methodologies such as lateral flow immunoassays and molecular-based assays, the authors suggest the blending of several tools: a paper-based result and common mobile devices. A patient at home could take a nasal swab and use a POC device to return a colorimetric result, which could then be captured by mobile phone and rapidly sent to a clinician for analysis. They conclude that such a method could "provide new insights into designing POC COVID-19 diagnostics and ultimately improve the health care system to combat this and similar diseases."
In this brief editorial article published in the journal Diagnostics, Tiawanese researchers Yang et al. provide their insights into a more convenient at-home point-of-care (POC) testing method for the COVID-19 illness. Noting the urgency of the associated pandemic and the various costs associated with traditional testing methodologies such as lateral flow immunoassays and molecular-based assays, the authors suggest the blending of several tools: a paper-based result and common mobile devices. A patient at home could take a nasal swab and use a POC device to return a colorimetric result, which could then be captured by mobile phone and rapidly sent to a clinician for analysis. They conclude that such a method could "provide new insights into designing POC COVID-19 diagnostics and ultimately improve the health care system to combat this and similar diseases."







