Posted on May 11, 2021
By LabLynx
Journal articles
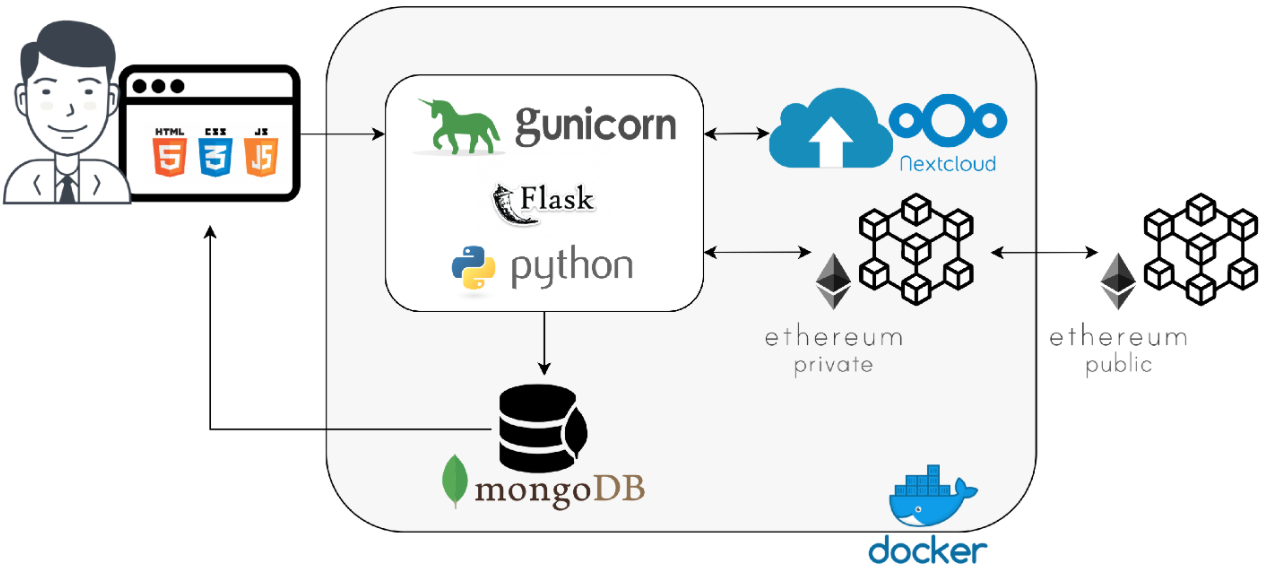
What do the internet of things (IoT), cloud computing, and blockchain all bring to the table of healthcare, particularly during a pandemic? Celesti
et al. demonstrate in this 2020 article published in
Sensors that the considerate combination of these technologies can lead to a "scenario where nurses, technicians, and medical doctors belonging to different hospitals cooperate through their federated hospital clouds to form a virtual health team able to carry out a healthcare workflow in secure fashion." The authors show how an IoT-connected laboratory that feeds its instrument data into a federated hospital cloud integrated with a blockchain engine can lead to less patient movement and better security of patient data, while also allowing nurses, doctors, and other practitioners from any of the connected hospitals to review results and issue treatments from a distance. They show the results of several experiments with such a system and conclude by promoting its benefits, as well as the possibility of extending the system to the pharmacy world.
Posted on May 3, 2021
By LabLynx
Journal articles
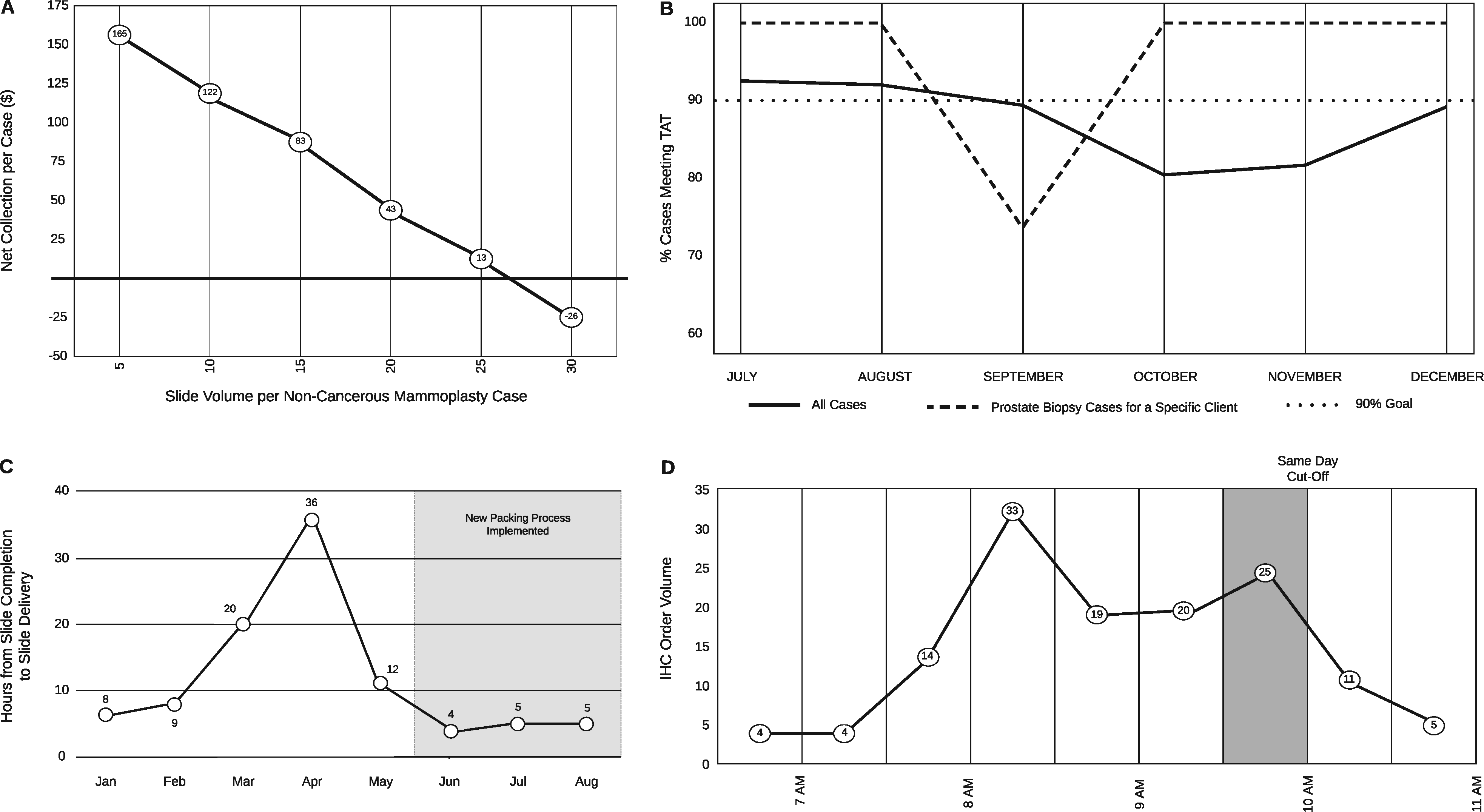
In this brief 2020 paper published in
JAMIA Open, Seifert
et al. of the University of Florida health system share their experiences with taking their Beaker LIS implementation and improving its data tracking capabilities, which in turn improved "trainee education, slide logistics, staffing and instrumentation lobbying, and task tracking" within their anatomic pathology laboratories. After a brief introduction, the authors demonstrate Beaker's status board and weaknesses, and then show how they used Beaker's MyReports module to develop an improved status board. They then state six significant challenges and how they used the adapted LIS tools to solve them. They conclude that "the technical and/or functionality framework that we demonstrated in this manuscript could be adapted by other institutions to address common problems encountered by anatomic pathology laboratories."
Posted on April 27, 2021
By LabLynx
Journal articles

In this 2021 paper published in
Frontiers in Chemistry, Myers
et al. of Restek and Verity Analytics provide details of their research on better terpene analysis methods in
Cannabis plant materials. The research focused on " the more traditional headspace syringe (HS syringe) and liquid injection syringe (LI syringe) approaches" compared to two more modern approaches to terpene analysis: HS-solid-phase microextraction Arrow (HS-SPME Arrow) and DI-SPME Arrow. After describing their experiments and results, the authors made multiple conclusions. First, they found "that DI-SPME Arrow performed better than HS-SPME Arrow; however, both of these approaches outperformed HS syringe for the extraction and analyses of terpenes." However, their results that LI syringe proved best, though not without some considerations concerning instrument uptime, compared to other methods. They also make a number of recommendations to cannabis laboratories and scientific researchers based upon their findings in order to "improve the science of cannabis testing."
Posted on April 21, 2021
By LabLynx
Journal articles
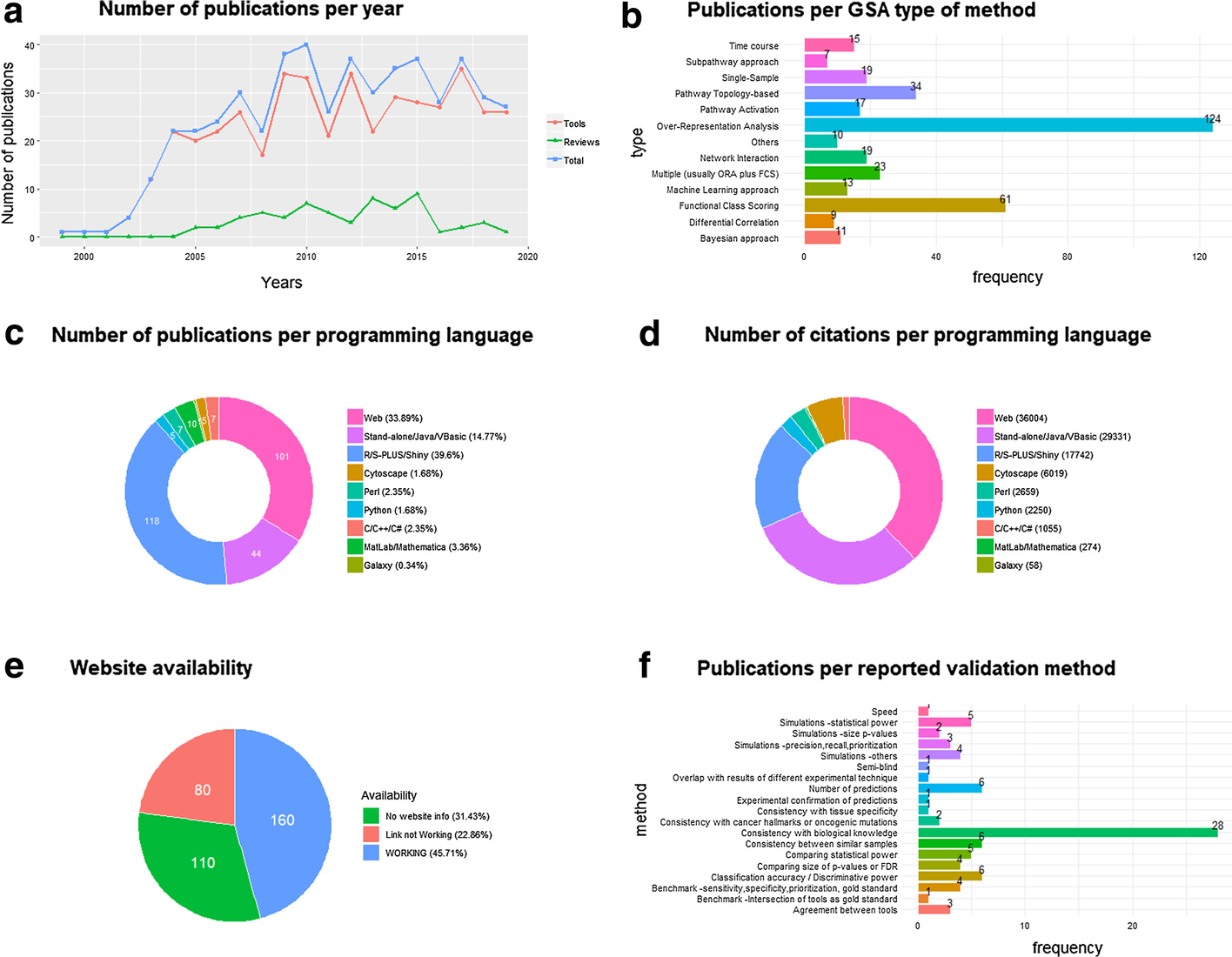
In this 2021 paper published in
BMC Bioinformatics, Xie
et al. examine big omics and gene set analysis, a common analytical technique for bioinformaticists. The authors argue that biomedical science researchers have not been using the best GSA methods among those that are actually available. They chose to conduct research of popularity and performance based on prior described validation strategies and existing benchmark studies. Noting that popular methods are not always the best methods, the authors conclude with five points from their research, including a need for deeper discussions and research about GSA methods and more rigorous validation procedures for GSA tools. They also provide examples of evaluational tools for GSA and other bioinformatics software.
Posted on April 12, 2021
By LabLynx
Journal articles
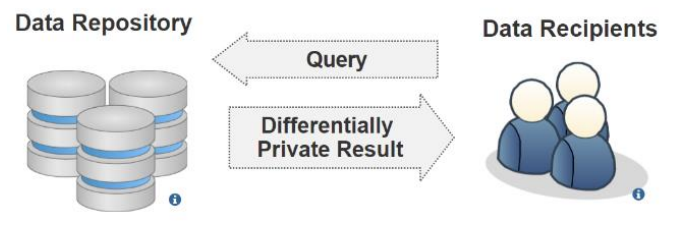
Last week's article looked at inference attacks in the cloud, but they happen in other ways as well. In this 2021 paper, Chong reviews the available literature on inference and other privacy attacks that occur on published healthcare information originally sourced from electronic health records (EHRs) and other health informatics systems. Ethical sharing of this kind of data with researchers conducting statistical analysis, improving clinical decision making, etc. is important, but it must preserve the privacy of the underlying individuals. After briefly discussing data publishing with a strong privacy focus, Chong looks at how healthcare data is typically stored and what threats exist against such data. The author then examines two well-established privacy models, including their strengths and limitations, that can be used to limit those threats: data anonymization and differential privacy. Despite these models, Chong closes by noting that "preserving privacy in healthcare data still poses several unsolved privacy and utility challenges" and expounds on areas for future research into those challenges.
Posted on April 7, 2021
By LabLynx
Journal articles
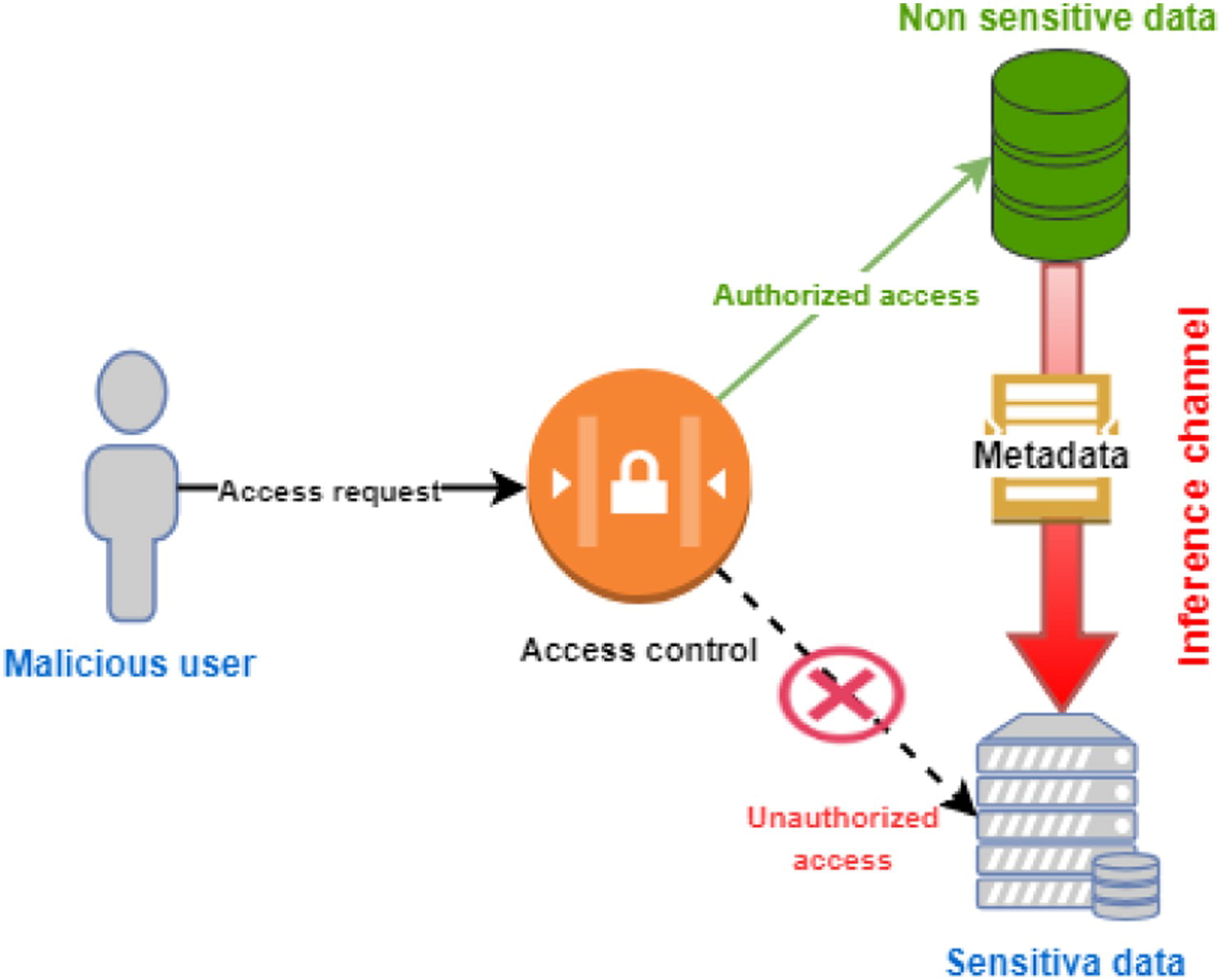
Inference attacks occur when a (typically) malicious actor is able to infer from public information more sensitive information about a database or individual without directly accessing the original database or sensitive user information. That these sorts of attacks can be successful against applications and data on cloud-based infrastructure should not be a surprise to seasoned IT veterans. Recognizing these concerns, Jebali
et al. examine the state of inference attacks in the cloud, the research that has been conducted so far on such attacks, and what can be done about them. Aside from examining a variety of security concerns in both communicating and non-communicating servers, the authors also address the inference problem, offering a potential solution that will in theory "optimize data distribution without the need to query the workload, then partition the database in the cloud by taking into consideration access control policies and data utility, before finally running a query evaluation model on a big data framework to securely process distributed queries while retaining access control." They conclude by recognizing more research must be done to test their proposed solution, and noting that past research has often overlooked other types of inference sources such as inclusion dependencies, join dependencies, and multivalued dependencies, which should be further examined.
Posted on March 30, 2021
By LabLynx
Journal articles
In this 2020 paper published in
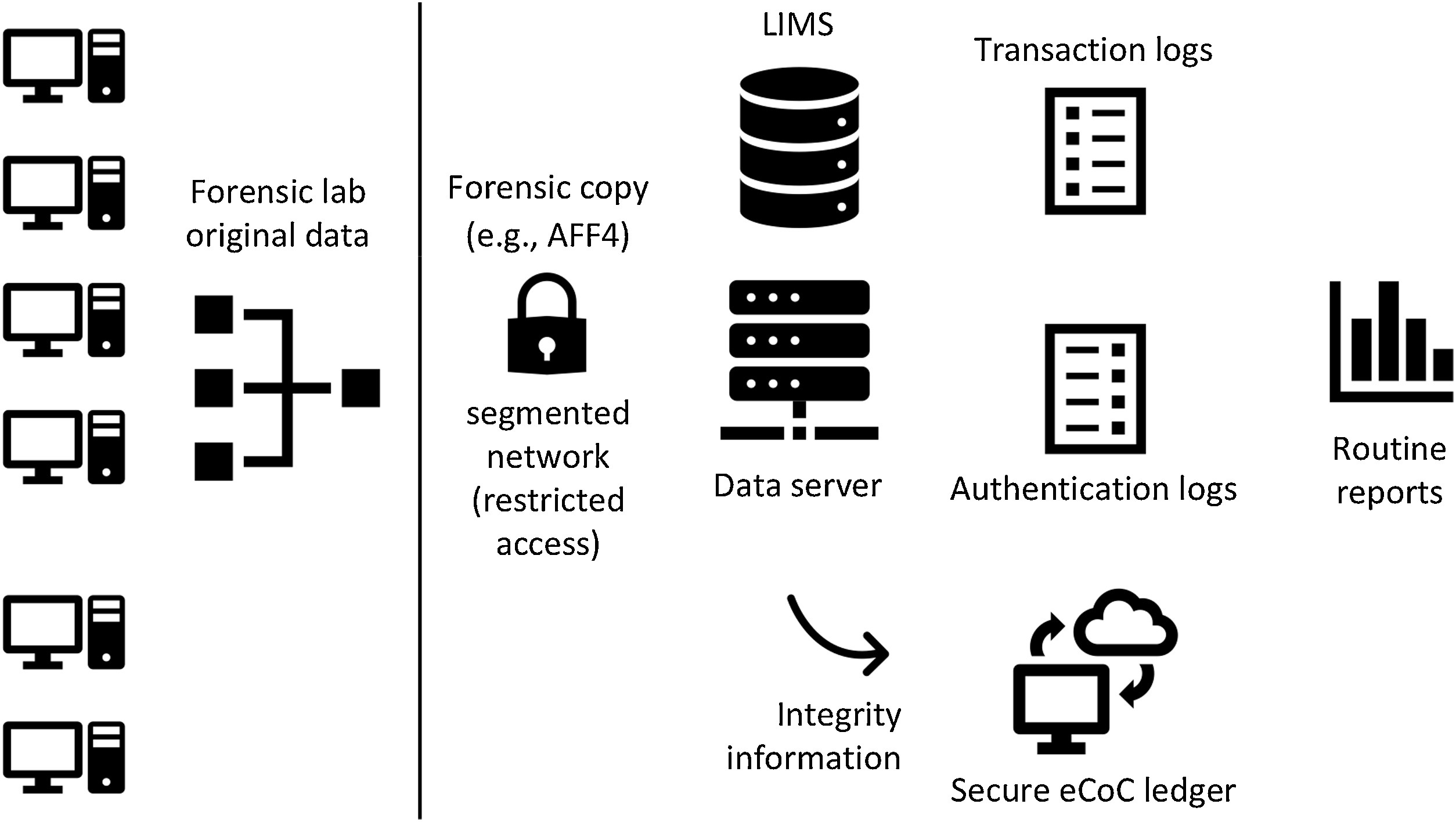
Forensic Science International, Casey and Souvignet present their ideas on how forensic laboratories with significant digital footprints should best prepare their operations to ensure their processes and digital data can be independently verified. Their recommendations come as they recognize how "digital transformations can undermine the core principles and processes of forensic laboratories" when those transformations are not planned and implemented well, risking the lab's integrity as well as the fundamental rights of individuals being forensically examined. After a brief introduction, the authors then present five risk scenarios and four technological improvement scenarios, followed by a discussion of a series of risk management practices to ensure digital transformations in the forensic laboratory are optimized and effective. They finally touch upon the value of in-house expertise and quality assurance practices in the lab, before concluding that "with proper forethought and preparation, forensic laboratories can employ technology and advanced data analytics to enhance existing services and create new services, while respecting fundamental human rights."
Posted on March 22, 2021
By LabLynx
Journal articles
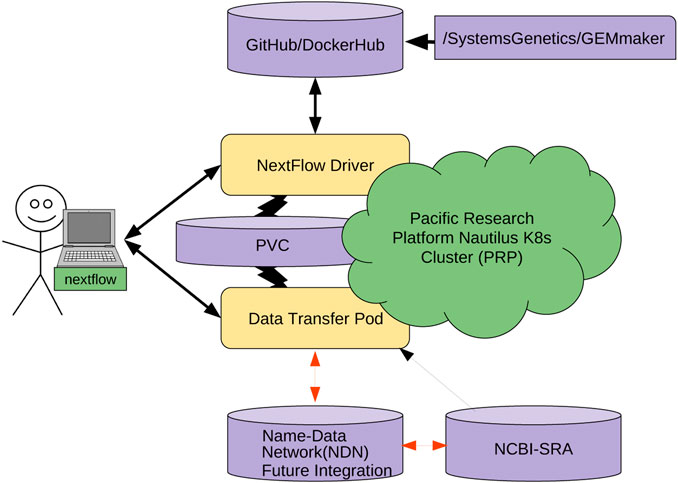
Managing and sharing big data in the cloud has its own set of cyberinfrastructure challenges, and that includes genomic research data. With large data sets being stored in numerous geographically distributed repositories around the world, effectively using this data for research can become enormously difficult. As such, Ogle
et al., writing in
Frontiers in Big Data, present their efforts towards reducing these network challenges using an internet architecture called named data networking (NDN). After discussing NDN, the authors describe the problems that come from wanting to manage and use big genomic data sets, as well as how NDN-based solutions can alleviate those problems. They then describe their method towards implementing NDN with genomics workflow tool GEMmaker in a cloud computing platform called the Pacific Research Platform. Through their efforts, they conclude that "NDN can serve data from anywhere, simplifying data management," and when integrated with GEMmaker, "in-network caching can speed up data retrieval and insertion into the workflow by six times," further improving use of big genomic data in practical research.
Posted on March 15, 2021
By LabLynx
Journal articles
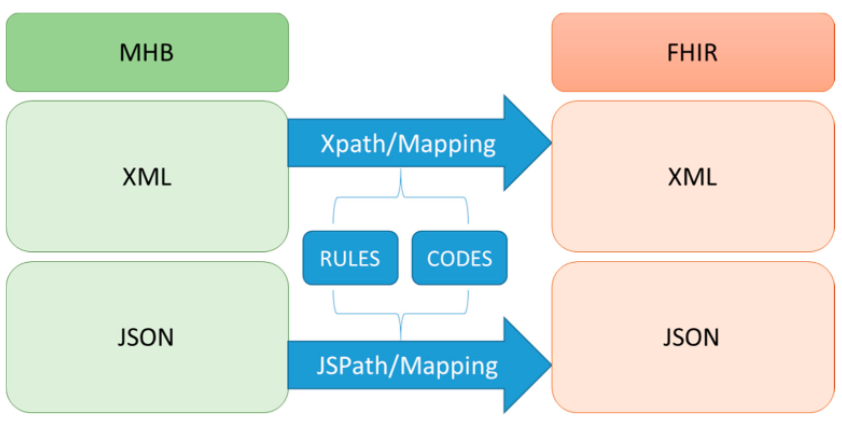
In this 2020 paper published in the journal
Sustainability, Lee
et al. provide details on their FHIR-based personal health record (PHR) system in Taiwan, which arguably has one of the best health systems in the world. With the Taiwan National Health Insurance Administration wanting to give its citizens more control over the health data, the demand has been there to improve how the average person can take a more proactive stance with their medical treatment. After reviewing relevant literature, the authors describe the evolution of their PHR platform, including content analysis, data conversion, networking, authentication, and security. Then, based upon their results, they conclude that the process of taking PHR data and applying international standards to its organization and presentation, they were able to effectively "achieve interoperability as well as capture and share data using FHIR resources and formats" while maintaining a sturdy but flexible user authentication mechanism.
Posted on March 8, 2021
By LabLynx
Journal articles
A data management plan (DMP) is a formally developed document that describes how data shall be handled both during research efforts and afterwards. However, as Weng and Thoben point out in this 2020 paper, "it a problem that DMPs [typically] do not include a structured approach for the identification or mitigation of risks" to that research data. The authors develop and propose a generic risk catalog for researchers to use in the development of their DMP, while demonstrating that catalog's value. After providing and introduction and related work, they present their catalog, developed from interviews with researchers across multiple disciplines. They then compare that catalog to 13 DMPs published in the journal
RIO. They conclude that their approach is "useful for identifying general risks in DMPs" and that it may also be useful to research "funders, since it makes it possible for them to evaluate basic investment risks of proposed projects."
Posted on March 1, 2021
By LabLynx
Journal articles
Cannabis-related businesses (CRBs) in U.S. states where cannabis is legal have long had difficulties finding the banking and financial services they require in a market where the federal government continues to consider cannabis illegal. Working in cash-only situations while trying to keep careful accounting, all while trying to keep the business afloat and clear of legal troubles, remains a challenge. This late 2020 paper by Owens-Ott recognizes that challenge and examines it further, delving into the world of the certified public accountant (CPA) and the role—or lack thereof—they are willing to play working with CRBs. The author conducted a qualitative study asking 1. why some CPAs are unwilling to provide services to CRBs, 2. how CRBs compensate for the lack of those services, and 3. what CPAs need to know in order to provide effective services to CRBs. Owens-Ott concludes that "competent and knowledgeable CPAs" willing to work with CRBs are out there, but the CPAs should be carefully vetted. And interested CPAs "must commit to acquiring and maintaining substantial specialized knowledge related to Tax Code Section 280E, internal controls for a cash-only or cash-intensive business, and the workings of the cannabis industry under the current regulatory conditions."
Posted on February 24, 2021
By LabLynx
Journal articles
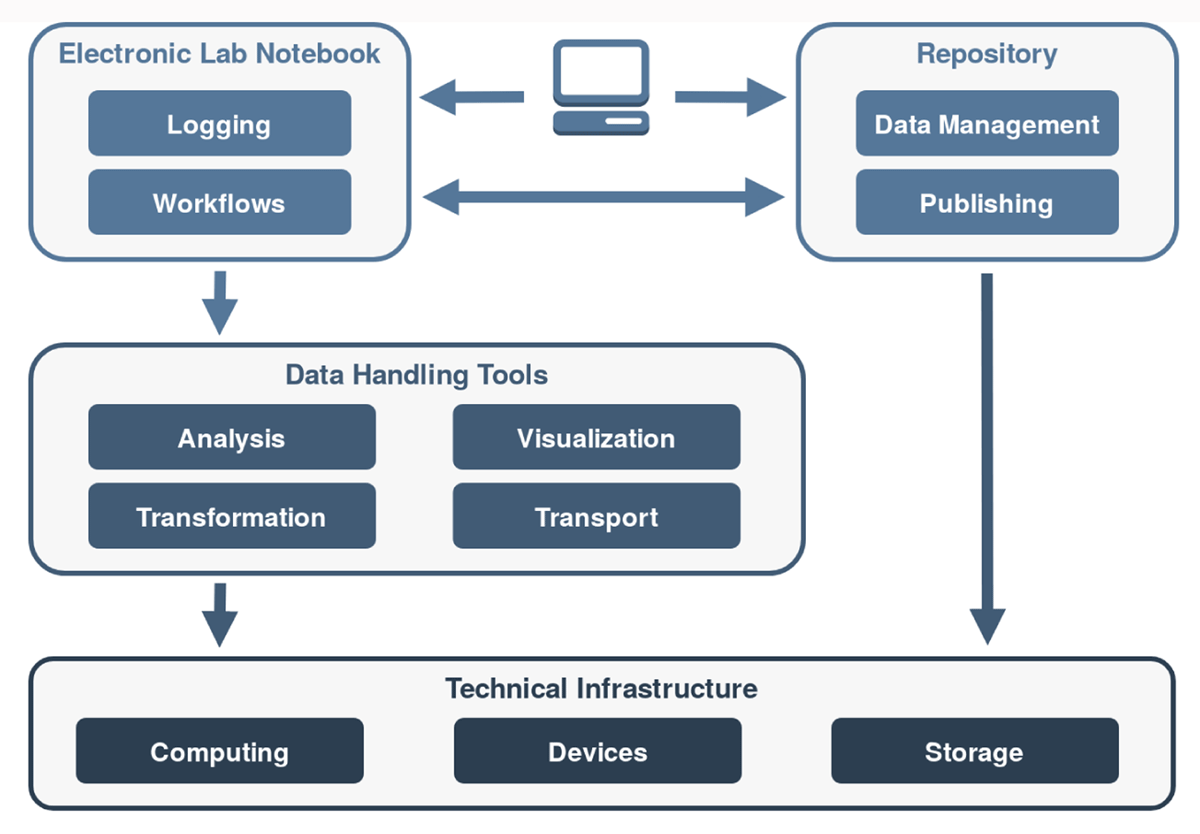
This 2021 paper published in
Data Science Journal explains Kadi4Mat, a combination electronic laboratory notebook (ELN) and data repository for better managing research and analyses in materials sciences. Noting the difficulty of extracting and using data obtained from experiments and simulations in the materials sciences, as well as the increasing levels of complexity in those experiments and simulations, Brandt
et al. sought to develop a comprehensive informatics solution capable of handling structured data storage and exchange in combination with documented and reproducible data analysis and visualization. Noting a wealth of open-source components for such an informatics project, the team provides a conceptual overview of the system and then dig into the details of its development using various open-source components. They then provide screenshots of records and data (and metadata) management, as well as workflow management. The authors conclude their solution helps manage "heterogeneous use cases of materials science disciplines," while being extensible enough to extend "the research data infrastructure to other disciplines" in the future.
Posted on February 15, 2021
By LabLynx
Journal articles
This guide by laboratory automation veteran Joe Liscouski takes an updated look at the state of implementing laboratory automation, with a strong emphasis on the importance of planning and management as early as possible. It also addresses the need for laboratory systems engineers (LSEs)—an update from laboratory automation engineers (LAEs)—and the institutional push required to develop educational opportunities for those LSEs. After looking at labs then and now, the author examines the transition to more automated labs, followed by seven detailed goals required for planning and managing that transition. He then closes by discussing LSEs and their education, concluding finally that "laboratory technology challenges can be addressed through planning activities using a series of goals," and that the lab of the future will implement those goals as early as possible, with the help of well-qualified LSEs fluent in both laboratory science and IT.
Posted on February 8, 2021
By LabLynx
Journal articles
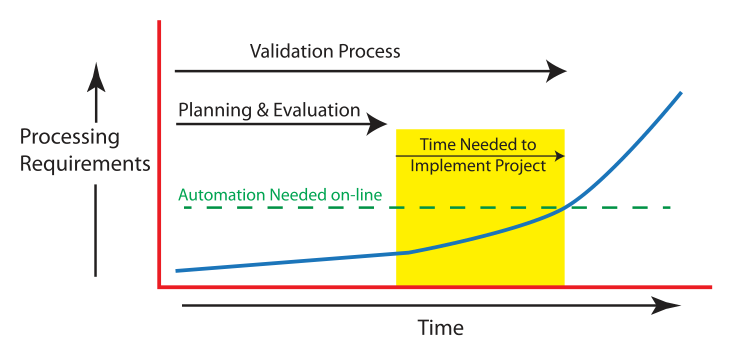
This white paper by laboratory automation veteran Joe Liscouski examines what it takes to fully realize a fully integrated laboratory informatics infrastructure, a strong pivot for many labs. He begins by discussing the worries and concerns of stakeholders about automation job losses, how AI and machine learning can play a role, and what equipment may be used, He then goes into lengthy discussion about how to specifically plan for automation, and at what level (no, partial, or full automation), as well as what it might cost. The author wraps up with a brief look at the steps required for actual project management. The conclusion? "Just as the manufacturing industries transitioned from cottage industries to production lines and then to integrated production-information systems, the execution of laboratory science has to tread a similar path if the demands for laboratory results are going to be met in a financially responsible manner." This makes "laboratory automation engineering" and planning a critical component of laboratories from the start.
Posted on February 1, 2021
By LabLynx
Journal articles
In this brief journal article published in
Data Intelligence, Weigel
et al. make their case for improving data object findability in order to benefit data and workflow management procedures for researchers. They argue that even with findable, accessible, interoperable, and reusable (FAIR) principles getting more attention, findability is the the starting point before anything else can be done, and that "support at the data infrastructure level for better automation of the processes dealing with data and workflows" is required. They succinctly describe the essential requirements for automating those processes, and then provide the basic building blocks that make up a potential solution, including—in the long-term—the application of machine learning. They conclude that adding automation to improve findability means that "researchers producing data can spend less time on data management and documentation, researchers reusing data and workflows will have access to metadata on a wider range of objects, and research administrators and funders may benefit from deeper insight into the impact of data-generating workflows."
Posted on January 25, 2021
By LabLynx
Journal articles
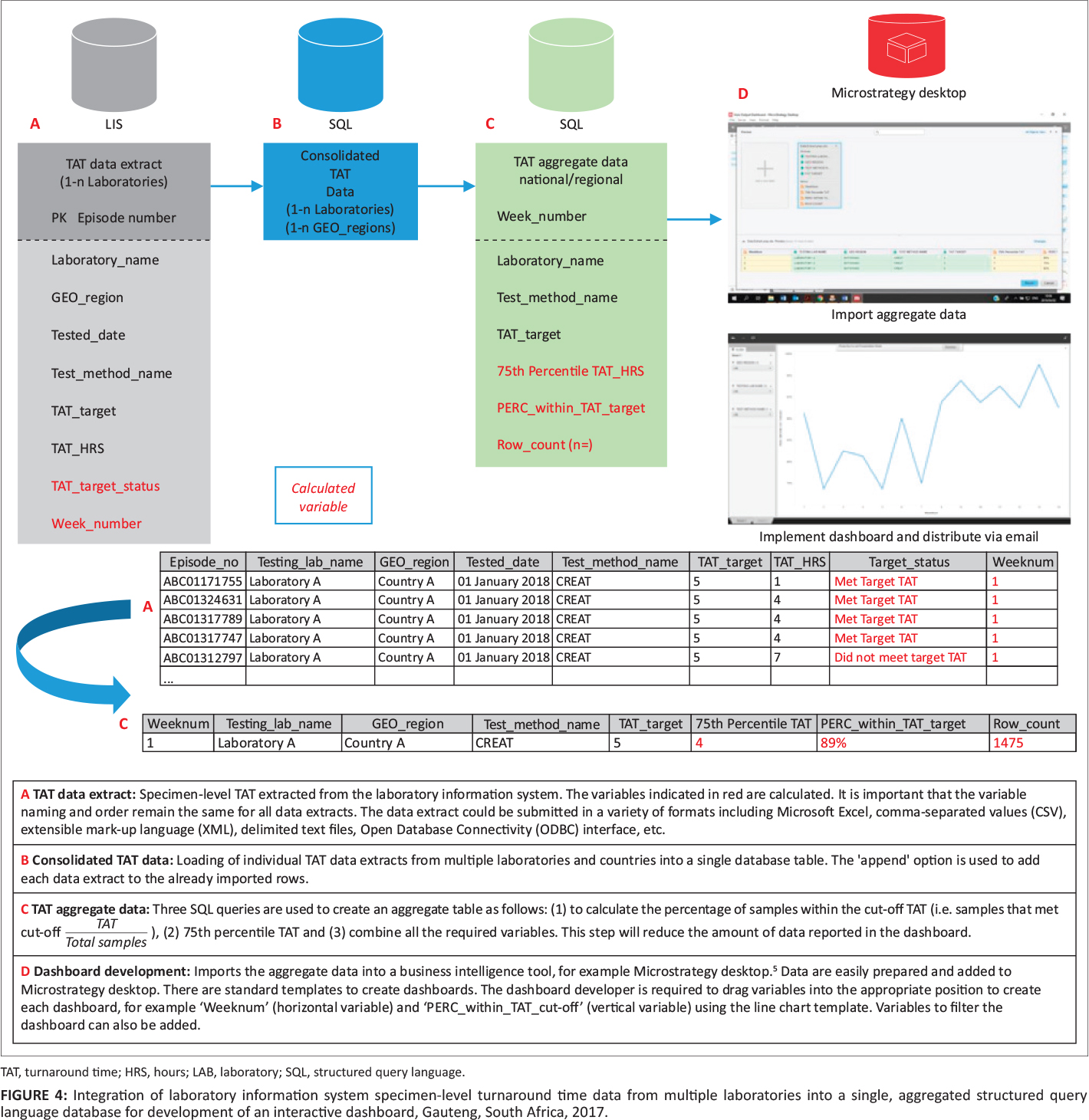
This companion piece to last week's article on laboratory turnaround time (TAT), sees Cassim
et al. take their experience developing and implementing a system for tracking TAT in a high-volume laboratory and assesses the actual impact such a system has. Using a retrospective study design and root cause analyses, the group looked at TAT outcomes over 122 weeks from a busy clinical pathology laboratory in a hospital in South Africa. After describing their methodology and results, the authors discussed the nuances of their results, including significant lessons learned. They conclude that not only is monitoring TAT to ensure timely reporting of results important, but also "vertical audits" of the results help identify bottlenecks for correction. They also emphasize "the importance of documenting and following through on corrective actions" associated with both those audits and the related quality management system in place at the lab.
Posted on January 18, 2021
By LabLynx
Journal articles
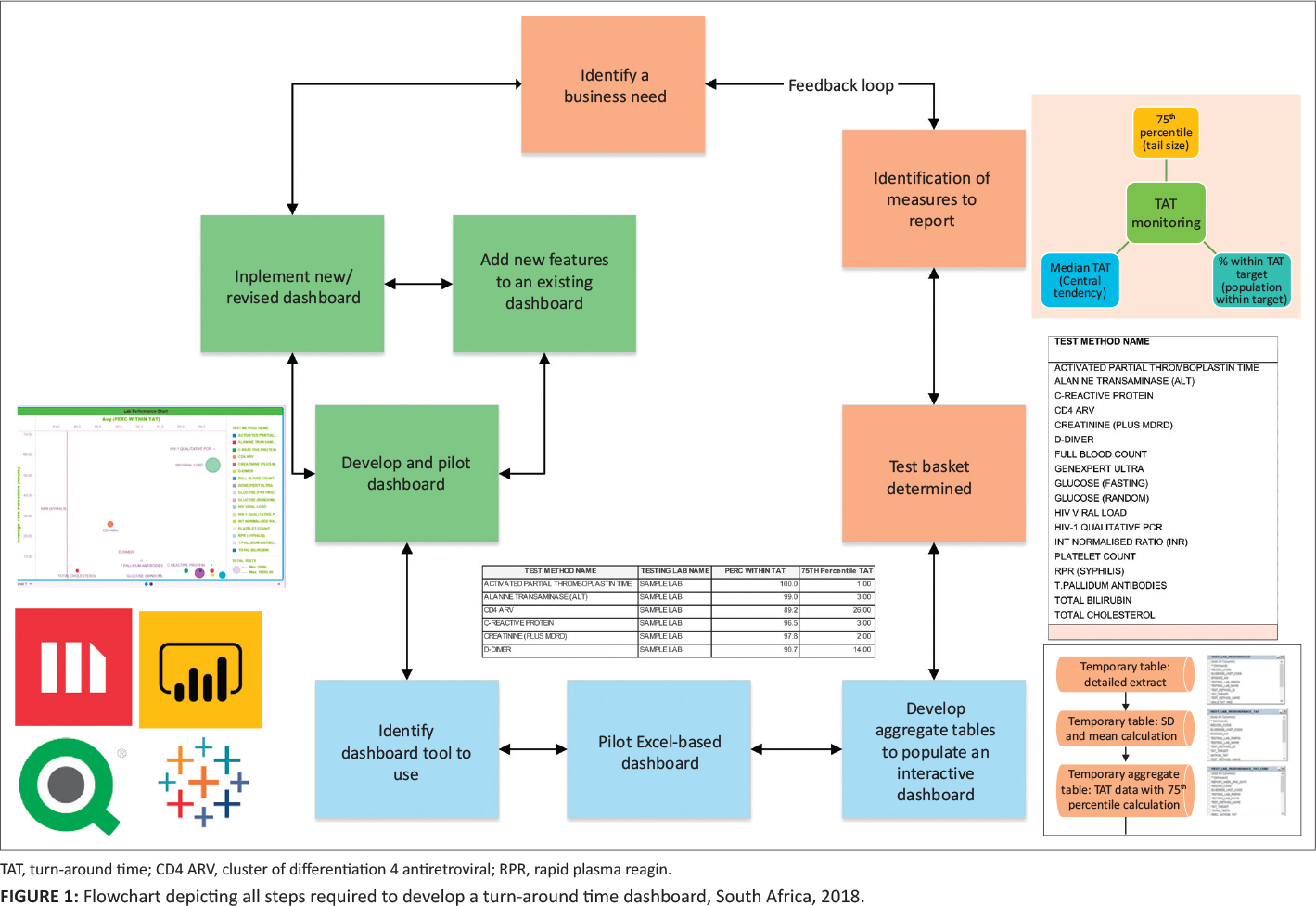
In this 2020 article published in the
African Journal of Laboratory Medicine, Cassim
et al. describe their experience developing and implementing a system for tracking turnaround time (TAT) in a high-volume laboratory. Seeking to have weekly reports of turnaround time—to better react to changes and improve policy—the authors developed a dashboard-based system for the South African National Health Laboratory Service and tested it in one if their higher-volume labs to assess the TAT monitoring system's performance. They conclude that while their dashboard "enables presentation of weekly TAT data to relevant business and laboratory managers, as part of the overall quality management portfolio of the organization," the actual act of "providing tools to assess TAT performance does not in itself imply corrective action or improvement." They emphasize that training on the system, as well as system performance measurements, gradual quality improvements, and the encouragement of a leadership-promoted business culture that supports the use of such data, are all required for the success of such a tool to be ensured.
Posted on January 11, 2021
By LabLynx
Journal articles
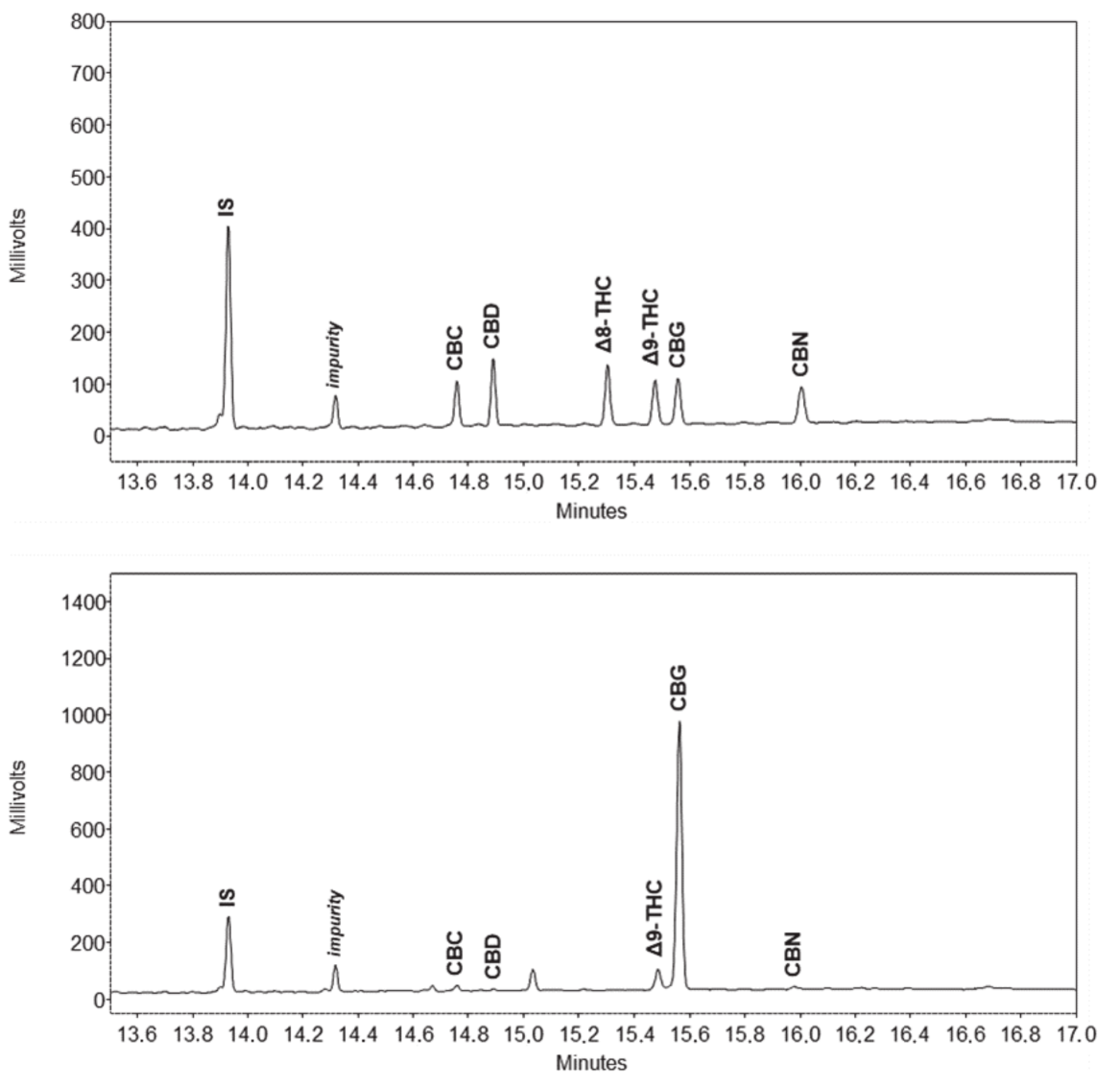
Over the years, a number of methods for detecting cannabinoids and terpenes in
Cannabis plant and related samples have been developed, recognizing the base problem of decarboxylation in gas chromatography. Some methods have proved better than others for cannabinoids, though typically differing methods are used for terpenes. However, could a method clearly detect both at the same time? Zekič and Križman demonstrate their gas chromatography method doing just that in this 2020 article published in
Molecules. After introducing their problem, the duo present their method, as an attempt "to find appropriate conditions, mainly in terms of sample preparation, for a simultaneous analysis of both groups of compounds, while keeping the overall experimental and instrumental setups simple." After discussing the method details and results, they conclude that their gas chromatography–flame ionization detection (GC-FID) method "provides a robust tool for simultaneous quantitative analysis of these two chemically different groups of analytes."
Posted on January 5, 2021
By LabLynx
Journal articles
In this 2020 paper published in
Developments in the Built Environment, Hartmann and Trappey share their years of experience working within the field of advanced engineering informatics, the pairing of "adequate computational tools" with modern engineering work, with the goal of improving the management of and collaboration related to increasingly complex engineering projects. While the duo don't directly address the specifics of such computational tools, they acknowledge engineering work as "knowledge-intensive" and detail the importance of formalizing that knowledge and its representation within various computational systems. They discuss the importance of knowledge representation and formation, as it informs research into advanced engineering informatics, and provide four practical examples provides by other researchers in the field. They then discuss the methodological approaches of implementing those more philosophical aspects towards developing knowledge representations, as well as how they should be verified and validated. They conclude that "knowledge representation is the main research effort that is required to develop technologies that not only automate mundane engineering tasks, but also provide engineers with tools that will allow them to do things they were not able to do before."
Posted on December 29, 2020
By LabLynx
Journal articles
Artificial intelligence (AI) is increasingly making its way into many aspects of health informatics, backed by a vision of improving healthcare outcomes and lowering healthcare costs. However, like many other such technologies, it comes with legal, ethical, and societal questions that deserve further exploration. Amann
et al. do just that in this 2020 paper published in
BMC Medical Informatics and Decision Making, examining the concept of "explainability," or why the AI came to the conclusion that it did in its task. The authors provide a brief amount of background before then examining AI-based clinical decision support systems (CDDSs) to provide various perspectives on the value of explainability in AI. They examine what explainability is from the perspective of technological perspective, then examine the legal, medical, and patient perspectives of explainability's importance. Finally, they examine the ethical implications of explainability using Beauchamp and Childress'
Principles of Biomedical Ethics. The authors conclude "that omitting explainability in CDDSs poses a threat to core ethical values in medicine and may have detrimental consequences for individual and public health."
 What do the internet of things (IoT), cloud computing, and blockchain all bring to the table of healthcare, particularly during a pandemic? Celesti et al. demonstrate in this 2020 article published in Sensors that the considerate combination of these technologies can lead to a "scenario where nurses, technicians, and medical doctors belonging to different hospitals cooperate through their federated hospital clouds to form a virtual health team able to carry out a healthcare workflow in secure fashion." The authors show how an IoT-connected laboratory that feeds its instrument data into a federated hospital cloud integrated with a blockchain engine can lead to less patient movement and better security of patient data, while also allowing nurses, doctors, and other practitioners from any of the connected hospitals to review results and issue treatments from a distance. They show the results of several experiments with such a system and conclude by promoting its benefits, as well as the possibility of extending the system to the pharmacy world.
What do the internet of things (IoT), cloud computing, and blockchain all bring to the table of healthcare, particularly during a pandemic? Celesti et al. demonstrate in this 2020 article published in Sensors that the considerate combination of these technologies can lead to a "scenario where nurses, technicians, and medical doctors belonging to different hospitals cooperate through their federated hospital clouds to form a virtual health team able to carry out a healthcare workflow in secure fashion." The authors show how an IoT-connected laboratory that feeds its instrument data into a federated hospital cloud integrated with a blockchain engine can lead to less patient movement and better security of patient data, while also allowing nurses, doctors, and other practitioners from any of the connected hospitals to review results and issue treatments from a distance. They show the results of several experiments with such a system and conclude by promoting its benefits, as well as the possibility of extending the system to the pharmacy world.
 In this brief 2020 paper published in JAMIA Open, Seifert et al. of the University of Florida health system share their experiences with taking their Beaker LIS implementation and improving its data tracking capabilities, which in turn improved "trainee education, slide logistics, staffing and instrumentation lobbying, and task tracking" within their anatomic pathology laboratories. After a brief introduction, the authors demonstrate Beaker's status board and weaknesses, and then show how they used Beaker's MyReports module to develop an improved status board. They then state six significant challenges and how they used the adapted LIS tools to solve them. They conclude that "the technical and/or functionality framework that we demonstrated in this manuscript could be adapted by other institutions to address common problems encountered by anatomic pathology laboratories."
In this brief 2020 paper published in JAMIA Open, Seifert et al. of the University of Florida health system share their experiences with taking their Beaker LIS implementation and improving its data tracking capabilities, which in turn improved "trainee education, slide logistics, staffing and instrumentation lobbying, and task tracking" within their anatomic pathology laboratories. After a brief introduction, the authors demonstrate Beaker's status board and weaknesses, and then show how they used Beaker's MyReports module to develop an improved status board. They then state six significant challenges and how they used the adapted LIS tools to solve them. They conclude that "the technical and/or functionality framework that we demonstrated in this manuscript could be adapted by other institutions to address common problems encountered by anatomic pathology laboratories."
 In this 2021 paper published in Frontiers in Chemistry, Myers et al. of Restek and Verity Analytics provide details of their research on better terpene analysis methods in Cannabis plant materials. The research focused on " the more traditional headspace syringe (HS syringe) and liquid injection syringe (LI syringe) approaches" compared to two more modern approaches to terpene analysis: HS-solid-phase microextraction Arrow (HS-SPME Arrow) and DI-SPME Arrow. After describing their experiments and results, the authors made multiple conclusions. First, they found "that DI-SPME Arrow performed better than HS-SPME Arrow; however, both of these approaches outperformed HS syringe for the extraction and analyses of terpenes." However, their results that LI syringe proved best, though not without some considerations concerning instrument uptime, compared to other methods. They also make a number of recommendations to cannabis laboratories and scientific researchers based upon their findings in order to "improve the science of cannabis testing."
In this 2021 paper published in Frontiers in Chemistry, Myers et al. of Restek and Verity Analytics provide details of their research on better terpene analysis methods in Cannabis plant materials. The research focused on " the more traditional headspace syringe (HS syringe) and liquid injection syringe (LI syringe) approaches" compared to two more modern approaches to terpene analysis: HS-solid-phase microextraction Arrow (HS-SPME Arrow) and DI-SPME Arrow. After describing their experiments and results, the authors made multiple conclusions. First, they found "that DI-SPME Arrow performed better than HS-SPME Arrow; however, both of these approaches outperformed HS syringe for the extraction and analyses of terpenes." However, their results that LI syringe proved best, though not without some considerations concerning instrument uptime, compared to other methods. They also make a number of recommendations to cannabis laboratories and scientific researchers based upon their findings in order to "improve the science of cannabis testing."
 In this 2021 paper published in BMC Bioinformatics, Xie et al. examine big omics and gene set analysis, a common analytical technique for bioinformaticists. The authors argue that biomedical science researchers have not been using the best GSA methods among those that are actually available. They chose to conduct research of popularity and performance based on prior described validation strategies and existing benchmark studies. Noting that popular methods are not always the best methods, the authors conclude with five points from their research, including a need for deeper discussions and research about GSA methods and more rigorous validation procedures for GSA tools. They also provide examples of evaluational tools for GSA and other bioinformatics software.
In this 2021 paper published in BMC Bioinformatics, Xie et al. examine big omics and gene set analysis, a common analytical technique for bioinformaticists. The authors argue that biomedical science researchers have not been using the best GSA methods among those that are actually available. They chose to conduct research of popularity and performance based on prior described validation strategies and existing benchmark studies. Noting that popular methods are not always the best methods, the authors conclude with five points from their research, including a need for deeper discussions and research about GSA methods and more rigorous validation procedures for GSA tools. They also provide examples of evaluational tools for GSA and other bioinformatics software.
 Last week's article looked at inference attacks in the cloud, but they happen in other ways as well. In this 2021 paper, Chong reviews the available literature on inference and other privacy attacks that occur on published healthcare information originally sourced from electronic health records (EHRs) and other health informatics systems. Ethical sharing of this kind of data with researchers conducting statistical analysis, improving clinical decision making, etc. is important, but it must preserve the privacy of the underlying individuals. After briefly discussing data publishing with a strong privacy focus, Chong looks at how healthcare data is typically stored and what threats exist against such data. The author then examines two well-established privacy models, including their strengths and limitations, that can be used to limit those threats: data anonymization and differential privacy. Despite these models, Chong closes by noting that "preserving privacy in healthcare data still poses several unsolved privacy and utility challenges" and expounds on areas for future research into those challenges.
Last week's article looked at inference attacks in the cloud, but they happen in other ways as well. In this 2021 paper, Chong reviews the available literature on inference and other privacy attacks that occur on published healthcare information originally sourced from electronic health records (EHRs) and other health informatics systems. Ethical sharing of this kind of data with researchers conducting statistical analysis, improving clinical decision making, etc. is important, but it must preserve the privacy of the underlying individuals. After briefly discussing data publishing with a strong privacy focus, Chong looks at how healthcare data is typically stored and what threats exist against such data. The author then examines two well-established privacy models, including their strengths and limitations, that can be used to limit those threats: data anonymization and differential privacy. Despite these models, Chong closes by noting that "preserving privacy in healthcare data still poses several unsolved privacy and utility challenges" and expounds on areas for future research into those challenges.
 Inference attacks occur when a (typically) malicious actor is able to infer from public information more sensitive information about a database or individual without directly accessing the original database or sensitive user information. That these sorts of attacks can be successful against applications and data on cloud-based infrastructure should not be a surprise to seasoned IT veterans. Recognizing these concerns, Jebali et al. examine the state of inference attacks in the cloud, the research that has been conducted so far on such attacks, and what can be done about them. Aside from examining a variety of security concerns in both communicating and non-communicating servers, the authors also address the inference problem, offering a potential solution that will in theory "optimize data distribution without the need to query the workload, then partition the database in the cloud by taking into consideration access control policies and data utility, before finally running a query evaluation model on a big data framework to securely process distributed queries while retaining access control." They conclude by recognizing more research must be done to test their proposed solution, and noting that past research has often overlooked other types of inference sources such as inclusion dependencies, join dependencies, and multivalued dependencies, which should be further examined.
Inference attacks occur when a (typically) malicious actor is able to infer from public information more sensitive information about a database or individual without directly accessing the original database or sensitive user information. That these sorts of attacks can be successful against applications and data on cloud-based infrastructure should not be a surprise to seasoned IT veterans. Recognizing these concerns, Jebali et al. examine the state of inference attacks in the cloud, the research that has been conducted so far on such attacks, and what can be done about them. Aside from examining a variety of security concerns in both communicating and non-communicating servers, the authors also address the inference problem, offering a potential solution that will in theory "optimize data distribution without the need to query the workload, then partition the database in the cloud by taking into consideration access control policies and data utility, before finally running a query evaluation model on a big data framework to securely process distributed queries while retaining access control." They conclude by recognizing more research must be done to test their proposed solution, and noting that past research has often overlooked other types of inference sources such as inclusion dependencies, join dependencies, and multivalued dependencies, which should be further examined.
 In this 2020 paper published in Forensic Science International, Casey and Souvignet present their ideas on how forensic laboratories with significant digital footprints should best prepare their operations to ensure their processes and digital data can be independently verified. Their recommendations come as they recognize how "digital transformations can undermine the core principles and processes of forensic laboratories" when those transformations are not planned and implemented well, risking the lab's integrity as well as the fundamental rights of individuals being forensically examined. After a brief introduction, the authors then present five risk scenarios and four technological improvement scenarios, followed by a discussion of a series of risk management practices to ensure digital transformations in the forensic laboratory are optimized and effective. They finally touch upon the value of in-house expertise and quality assurance practices in the lab, before concluding that "with proper forethought and preparation, forensic laboratories can employ technology and advanced data analytics to enhance existing services and create new services, while respecting fundamental human rights."
In this 2020 paper published in Forensic Science International, Casey and Souvignet present their ideas on how forensic laboratories with significant digital footprints should best prepare their operations to ensure their processes and digital data can be independently verified. Their recommendations come as they recognize how "digital transformations can undermine the core principles and processes of forensic laboratories" when those transformations are not planned and implemented well, risking the lab's integrity as well as the fundamental rights of individuals being forensically examined. After a brief introduction, the authors then present five risk scenarios and four technological improvement scenarios, followed by a discussion of a series of risk management practices to ensure digital transformations in the forensic laboratory are optimized and effective. They finally touch upon the value of in-house expertise and quality assurance practices in the lab, before concluding that "with proper forethought and preparation, forensic laboratories can employ technology and advanced data analytics to enhance existing services and create new services, while respecting fundamental human rights."
 Managing and sharing big data in the cloud has its own set of cyberinfrastructure challenges, and that includes genomic research data. With large data sets being stored in numerous geographically distributed repositories around the world, effectively using this data for research can become enormously difficult. As such, Ogle et al., writing in Frontiers in Big Data, present their efforts towards reducing these network challenges using an internet architecture called named data networking (NDN). After discussing NDN, the authors describe the problems that come from wanting to manage and use big genomic data sets, as well as how NDN-based solutions can alleviate those problems. They then describe their method towards implementing NDN with genomics workflow tool GEMmaker in a cloud computing platform called the Pacific Research Platform. Through their efforts, they conclude that "NDN can serve data from anywhere, simplifying data management," and when integrated with GEMmaker, "in-network caching can speed up data retrieval and insertion into the workflow by six times," further improving use of big genomic data in practical research.
Managing and sharing big data in the cloud has its own set of cyberinfrastructure challenges, and that includes genomic research data. With large data sets being stored in numerous geographically distributed repositories around the world, effectively using this data for research can become enormously difficult. As such, Ogle et al., writing in Frontiers in Big Data, present their efforts towards reducing these network challenges using an internet architecture called named data networking (NDN). After discussing NDN, the authors describe the problems that come from wanting to manage and use big genomic data sets, as well as how NDN-based solutions can alleviate those problems. They then describe their method towards implementing NDN with genomics workflow tool GEMmaker in a cloud computing platform called the Pacific Research Platform. Through their efforts, they conclude that "NDN can serve data from anywhere, simplifying data management," and when integrated with GEMmaker, "in-network caching can speed up data retrieval and insertion into the workflow by six times," further improving use of big genomic data in practical research.
 In this 2020 paper published in the journal Sustainability, Lee et al. provide details on their FHIR-based personal health record (PHR) system in Taiwan, which arguably has one of the best health systems in the world. With the Taiwan National Health Insurance Administration wanting to give its citizens more control over the health data, the demand has been there to improve how the average person can take a more proactive stance with their medical treatment. After reviewing relevant literature, the authors describe the evolution of their PHR platform, including content analysis, data conversion, networking, authentication, and security. Then, based upon their results, they conclude that the process of taking PHR data and applying international standards to its organization and presentation, they were able to effectively "achieve interoperability as well as capture and share data using FHIR resources and formats" while maintaining a sturdy but flexible user authentication mechanism.
In this 2020 paper published in the journal Sustainability, Lee et al. provide details on their FHIR-based personal health record (PHR) system in Taiwan, which arguably has one of the best health systems in the world. With the Taiwan National Health Insurance Administration wanting to give its citizens more control over the health data, the demand has been there to improve how the average person can take a more proactive stance with their medical treatment. After reviewing relevant literature, the authors describe the evolution of their PHR platform, including content analysis, data conversion, networking, authentication, and security. Then, based upon their results, they conclude that the process of taking PHR data and applying international standards to its organization and presentation, they were able to effectively "achieve interoperability as well as capture and share data using FHIR resources and formats" while maintaining a sturdy but flexible user authentication mechanism.
 This 2021 paper published in Data Science Journal explains Kadi4Mat, a combination electronic laboratory notebook (ELN) and data repository for better managing research and analyses in materials sciences. Noting the difficulty of extracting and using data obtained from experiments and simulations in the materials sciences, as well as the increasing levels of complexity in those experiments and simulations, Brandt et al. sought to develop a comprehensive informatics solution capable of handling structured data storage and exchange in combination with documented and reproducible data analysis and visualization. Noting a wealth of open-source components for such an informatics project, the team provides a conceptual overview of the system and then dig into the details of its development using various open-source components. They then provide screenshots of records and data (and metadata) management, as well as workflow management. The authors conclude their solution helps manage "heterogeneous use cases of materials science disciplines," while being extensible enough to extend "the research data infrastructure to other disciplines" in the future.
This 2021 paper published in Data Science Journal explains Kadi4Mat, a combination electronic laboratory notebook (ELN) and data repository for better managing research and analyses in materials sciences. Noting the difficulty of extracting and using data obtained from experiments and simulations in the materials sciences, as well as the increasing levels of complexity in those experiments and simulations, Brandt et al. sought to develop a comprehensive informatics solution capable of handling structured data storage and exchange in combination with documented and reproducible data analysis and visualization. Noting a wealth of open-source components for such an informatics project, the team provides a conceptual overview of the system and then dig into the details of its development using various open-source components. They then provide screenshots of records and data (and metadata) management, as well as workflow management. The authors conclude their solution helps manage "heterogeneous use cases of materials science disciplines," while being extensible enough to extend "the research data infrastructure to other disciplines" in the future.
 This guide by laboratory automation veteran Joe Liscouski takes an updated look at the state of implementing laboratory automation, with a strong emphasis on the importance of planning and management as early as possible. It also addresses the need for laboratory systems engineers (LSEs)—an update from laboratory automation engineers (LAEs)—and the institutional push required to develop educational opportunities for those LSEs. After looking at labs then and now, the author examines the transition to more automated labs, followed by seven detailed goals required for planning and managing that transition. He then closes by discussing LSEs and their education, concluding finally that "laboratory technology challenges can be addressed through planning activities using a series of goals," and that the lab of the future will implement those goals as early as possible, with the help of well-qualified LSEs fluent in both laboratory science and IT.
This guide by laboratory automation veteran Joe Liscouski takes an updated look at the state of implementing laboratory automation, with a strong emphasis on the importance of planning and management as early as possible. It also addresses the need for laboratory systems engineers (LSEs)—an update from laboratory automation engineers (LAEs)—and the institutional push required to develop educational opportunities for those LSEs. After looking at labs then and now, the author examines the transition to more automated labs, followed by seven detailed goals required for planning and managing that transition. He then closes by discussing LSEs and their education, concluding finally that "laboratory technology challenges can be addressed through planning activities using a series of goals," and that the lab of the future will implement those goals as early as possible, with the help of well-qualified LSEs fluent in both laboratory science and IT.
 This white paper by laboratory automation veteran Joe Liscouski examines what it takes to fully realize a fully integrated laboratory informatics infrastructure, a strong pivot for many labs. He begins by discussing the worries and concerns of stakeholders about automation job losses, how AI and machine learning can play a role, and what equipment may be used, He then goes into lengthy discussion about how to specifically plan for automation, and at what level (no, partial, or full automation), as well as what it might cost. The author wraps up with a brief look at the steps required for actual project management. The conclusion? "Just as the manufacturing industries transitioned from cottage industries to production lines and then to integrated production-information systems, the execution of laboratory science has to tread a similar path if the demands for laboratory results are going to be met in a financially responsible manner." This makes "laboratory automation engineering" and planning a critical component of laboratories from the start.
This white paper by laboratory automation veteran Joe Liscouski examines what it takes to fully realize a fully integrated laboratory informatics infrastructure, a strong pivot for many labs. He begins by discussing the worries and concerns of stakeholders about automation job losses, how AI and machine learning can play a role, and what equipment may be used, He then goes into lengthy discussion about how to specifically plan for automation, and at what level (no, partial, or full automation), as well as what it might cost. The author wraps up with a brief look at the steps required for actual project management. The conclusion? "Just as the manufacturing industries transitioned from cottage industries to production lines and then to integrated production-information systems, the execution of laboratory science has to tread a similar path if the demands for laboratory results are going to be met in a financially responsible manner." This makes "laboratory automation engineering" and planning a critical component of laboratories from the start.
 This companion piece to last week's article on laboratory turnaround time (TAT), sees Cassim et al. take their experience developing and implementing a system for tracking TAT in a high-volume laboratory and assesses the actual impact such a system has. Using a retrospective study design and root cause analyses, the group looked at TAT outcomes over 122 weeks from a busy clinical pathology laboratory in a hospital in South Africa. After describing their methodology and results, the authors discussed the nuances of their results, including significant lessons learned. They conclude that not only is monitoring TAT to ensure timely reporting of results important, but also "vertical audits" of the results help identify bottlenecks for correction. They also emphasize "the importance of documenting and following through on corrective actions" associated with both those audits and the related quality management system in place at the lab.
This companion piece to last week's article on laboratory turnaround time (TAT), sees Cassim et al. take their experience developing and implementing a system for tracking TAT in a high-volume laboratory and assesses the actual impact such a system has. Using a retrospective study design and root cause analyses, the group looked at TAT outcomes over 122 weeks from a busy clinical pathology laboratory in a hospital in South Africa. After describing their methodology and results, the authors discussed the nuances of their results, including significant lessons learned. They conclude that not only is monitoring TAT to ensure timely reporting of results important, but also "vertical audits" of the results help identify bottlenecks for correction. They also emphasize "the importance of documenting and following through on corrective actions" associated with both those audits and the related quality management system in place at the lab.
 In this 2020 article published in the African Journal of Laboratory Medicine, Cassim et al. describe their experience developing and implementing a system for tracking turnaround time (TAT) in a high-volume laboratory. Seeking to have weekly reports of turnaround time—to better react to changes and improve policy—the authors developed a dashboard-based system for the South African National Health Laboratory Service and tested it in one if their higher-volume labs to assess the TAT monitoring system's performance. They conclude that while their dashboard "enables presentation of weekly TAT data to relevant business and laboratory managers, as part of the overall quality management portfolio of the organization," the actual act of "providing tools to assess TAT performance does not in itself imply corrective action or improvement." They emphasize that training on the system, as well as system performance measurements, gradual quality improvements, and the encouragement of a leadership-promoted business culture that supports the use of such data, are all required for the success of such a tool to be ensured.
In this 2020 article published in the African Journal of Laboratory Medicine, Cassim et al. describe their experience developing and implementing a system for tracking turnaround time (TAT) in a high-volume laboratory. Seeking to have weekly reports of turnaround time—to better react to changes and improve policy—the authors developed a dashboard-based system for the South African National Health Laboratory Service and tested it in one if their higher-volume labs to assess the TAT monitoring system's performance. They conclude that while their dashboard "enables presentation of weekly TAT data to relevant business and laboratory managers, as part of the overall quality management portfolio of the organization," the actual act of "providing tools to assess TAT performance does not in itself imply corrective action or improvement." They emphasize that training on the system, as well as system performance measurements, gradual quality improvements, and the encouragement of a leadership-promoted business culture that supports the use of such data, are all required for the success of such a tool to be ensured.
 Over the years, a number of methods for detecting cannabinoids and terpenes in Cannabis plant and related samples have been developed, recognizing the base problem of decarboxylation in gas chromatography. Some methods have proved better than others for cannabinoids, though typically differing methods are used for terpenes. However, could a method clearly detect both at the same time? Zekič and Križman demonstrate their gas chromatography method doing just that in this 2020 article published in Molecules. After introducing their problem, the duo present their method, as an attempt "to find appropriate conditions, mainly in terms of sample preparation, for a simultaneous analysis of both groups of compounds, while keeping the overall experimental and instrumental setups simple." After discussing the method details and results, they conclude that their gas chromatography–flame ionization detection (GC-FID) method "provides a robust tool for simultaneous quantitative analysis of these two chemically different groups of analytes."
Over the years, a number of methods for detecting cannabinoids and terpenes in Cannabis plant and related samples have been developed, recognizing the base problem of decarboxylation in gas chromatography. Some methods have proved better than others for cannabinoids, though typically differing methods are used for terpenes. However, could a method clearly detect both at the same time? Zekič and Križman demonstrate their gas chromatography method doing just that in this 2020 article published in Molecules. After introducing their problem, the duo present their method, as an attempt "to find appropriate conditions, mainly in terms of sample preparation, for a simultaneous analysis of both groups of compounds, while keeping the overall experimental and instrumental setups simple." After discussing the method details and results, they conclude that their gas chromatography–flame ionization detection (GC-FID) method "provides a robust tool for simultaneous quantitative analysis of these two chemically different groups of analytes."







